Solid State Nuclear Magnetic Resonance ( IF 3.2 ) Pub Date : 2020-11-04 , DOI: 10.1016/j.ssnmr.2020.101699 Maxime Yon , Franck Fayon , Dominique Massiot , Vincent Sarou-Kanian
|
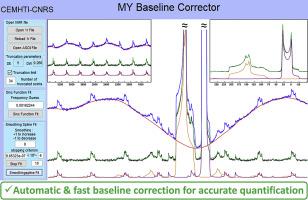
We present an algorithm suitable for automatically correcting rolling baseline coming from time-domain truncation induced by the dead time in pulse-acquire one-dimensional MAS NMR spectra. It relies on an iterative estimation of the baseline restricted in the time-domain by the dead time duration combined with a histogram filter allowing adaptive selection of the baseline points. This method does not make any assumption regarding the NMR resonances line shapes or widths and does not modify the acquired free induction decay points. This makes it suitable for accurate deconvolution and quantification of single-pulse MAS NMR spectra. The baseline correction accuracy is evaluated on synthetic solid-state spectra of 19F, 71Ga, and 23Na by comparing the fitted baseline to the theoretical one. The versatility of the algorithm is also exemplified on three additional solid-state spectra of 23Na and 71Ga. The algorithm is made available to the community through a user-friendly standalone Matlab® application.
中文翻译:

死区截断一维固态MAS NMR谱的迭代基线校正算法
我们提出了一种算法,该算法适用于自动校正来自脉冲获取一维MAS NMR谱中死区时间引起的时域截断的滚动基线。它依赖于死区持续时间在时域内受限的基线的迭代估计,并结合直方图过滤器,允许自适应选择基线点。该方法没有对NMR共振线的形状或宽度做任何假设,也没有修改获取的自由感应衰减点。这使其适用于单脉冲MAS NMR光谱的精确去卷积和定量。在19 F,71 Ga和23的合成固态光谱上评估基线校正精度通过将拟合的基线与理论基线进行比较,得出Na。该算法的多功能性还以23 Na和71 Ga的三个附加固态光谱为例。该算法可通过用户友好的独立Matlab®应用程序提供给社区。

























 京公网安备 11010802027423号
京公网安备 11010802027423号