当前位置:
X-MOL 学术
›
Chem. Soc. Rev.
›
论文详情
Our official English website, www.x-mol.net, welcomes your feedback! (Note: you will need to create a separate account there.)
Fundamental studies of functional nucleic acids: aptamers, riboswitches, ribozymes and DNAzymes.
Chemical Society Reviews ( IF 46.2 ) Pub Date : 2020-09-18 , DOI: 10.1039/d0cs00617c Ronald Micura 1 , Claudia Höbartner
Chemical Society Reviews ( IF 46.2 ) Pub Date : 2020-09-18 , DOI: 10.1039/d0cs00617c Ronald Micura 1 , Claudia Höbartner
Affiliation
|
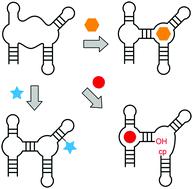
This review aims at juxtaposing common versus distinct structural and functional strategies that are applied by aptamers, riboswitches, and ribozymes/DNAzymes. Focusing on recently discovered systems, we begin our analysis with small-molecule binding aptamers, with emphasis on in vitro-selected fluorogenic RNA aptamers and their different modes of ligand binding and fluorescence activation. Fundamental insights are much needed to advance RNA imaging probes for detection of exo- and endogenous RNA and for RNA process tracking. Secondly, we discuss the latest gene expression–regulating mRNA riboswitches that respond to the alarmone ppGpp, to PRPP, to NAD+, to adenosine and cytidine diphosphates, and to precursors of thiamine biosynthesis (HMP-PP), and we outline new subclasses of SAM and tetrahydrofolate-binding RNA regulators. Many riboswitches bind protein enzyme cofactors that, in principle, can catalyse a chemical reaction. For RNA, however, only one system (glmS ribozyme) has been identified in Nature thus far that utilizes a small molecule – glucosamine-6-phosphate – to participate directly in reaction catalysis (phosphodiester cleavage). We wonder why that is the case and what is to be done to reveal such likely existing cellular activities that could be more diverse than currently imagined. Thirdly, this brings us to the four latest small nucleolytic ribozymes termed twister, twister-sister, pistol, and hatchet as well as to in vitro selected DNA and RNA enzymes that promote new chemistry, mainly by exploiting their ability for RNA labelling and nucleoside modification recognition. Enormous progress in understanding the strategies of nucleic acids catalysts has been made by providing thorough structural fundaments (e.g. first structure of a DNAzyme, structures of ribozyme transition state mimics) in combination with functional assays and atomic mutagenesis.
中文翻译:

功能核酸的基础研究:适体,核糖开关,核酶和脱氧核酶。
这篇综述的目的是将适体,核糖开关和核酶/ DNA酶应用的共同的和截然不同的结构和功能策略并列。着眼于最近发现的系统,我们从小分子结合适体开始分析,重点是体外选择的荧光RNA适体及其配体结合和荧光激活的不同模式。推进RNA成像探针用于检测外源和内源RNA以及进行RNA过程跟踪非常需要基础知识。其次,我们讨论了最新的基因表达调控mRNA的核糖开关,这些开关对警号ppGpp,PRPP,NAD +有反应,腺苷和胞苷二磷酸,以及硫胺素生物合成的前体(HMP-PP),我们概述了SAM和四氢叶酸结合RNA调节剂的新亚类。许多核糖开关结合蛋白质酶辅因子,原则上可以催化化学反应。然而,对于RNA,到目前为止,在自然界中仅发现了一种系统(glmS核酶),该系统利用小分子6-磷酸氨基葡萄糖来直接参与反应催化(磷酸二酯的裂解)。我们想知道为什么会这样,如何做才能揭示这种可能存在的细胞活动,其活动可能比目前想象的还要多样化。第三,这将我们带到了四个最新的小型核糖核酸酶中,它们分别被称为“扭转者”,“扭转者姐妹”,“手枪”和“柴刀”,以及在体外选择的DNA和RNA酶主要通过利用其RNA标记和核苷修饰识别能力来促进新化学。通过提供功能分析和原子诱变相结合的透彻结构基础(例如DNA酶的第一结构,核酶过渡态模拟物的结构),在理解核酸催化剂的策略方面取得了巨大进展。
更新日期:2020-10-19
中文翻译:

功能核酸的基础研究:适体,核糖开关,核酶和脱氧核酶。
这篇综述的目的是将适体,核糖开关和核酶/ DNA酶应用的共同的和截然不同的结构和功能策略并列。着眼于最近发现的系统,我们从小分子结合适体开始分析,重点是体外选择的荧光RNA适体及其配体结合和荧光激活的不同模式。推进RNA成像探针用于检测外源和内源RNA以及进行RNA过程跟踪非常需要基础知识。其次,我们讨论了最新的基因表达调控mRNA的核糖开关,这些开关对警号ppGpp,PRPP,NAD +有反应,腺苷和胞苷二磷酸,以及硫胺素生物合成的前体(HMP-PP),我们概述了SAM和四氢叶酸结合RNA调节剂的新亚类。许多核糖开关结合蛋白质酶辅因子,原则上可以催化化学反应。然而,对于RNA,到目前为止,在自然界中仅发现了一种系统(glmS核酶),该系统利用小分子6-磷酸氨基葡萄糖来直接参与反应催化(磷酸二酯的裂解)。我们想知道为什么会这样,如何做才能揭示这种可能存在的细胞活动,其活动可能比目前想象的还要多样化。第三,这将我们带到了四个最新的小型核糖核酸酶中,它们分别被称为“扭转者”,“扭转者姐妹”,“手枪”和“柴刀”,以及在体外选择的DNA和RNA酶主要通过利用其RNA标记和核苷修饰识别能力来促进新化学。通过提供功能分析和原子诱变相结合的透彻结构基础(例如DNA酶的第一结构,核酶过渡态模拟物的结构),在理解核酸催化剂的策略方面取得了巨大进展。

























 京公网安备 11010802027423号
京公网安备 11010802027423号