Abstract
Multi-protein (or enzyme) conjugates play a vital role in biosensing due to the integrated function of each component, such as biological recognition and signal amplification. In this work, a green self-assembled method for the synthesis of multi-functional protein-enzyme nanoflowers has been developed, in which no chemical modification and coupling reaction is needed to fabricate the fluorescent signal probe. The self-assembled protein-enzyme conjugates streptavidin (SA) -β-galactosidase (β-Gal)-CaHPO4 nanoflowers load sufficient enzymes without damaging their activity, which meets the requirements of signal tags for biosensing. Through integrated multi-function of biorecognition (SA) and signal amplification (β-Gal), the SA-β-Gal-CaHPO4 hybrid nanoflower-based fluorescent sensor exhibited an ultrasensitive detection of protein biomarker alpha-fetoprotein (AFP), with limits of detection at the fM level. The presented self-assembled strategy can be extensively applied to develop on-demand protein-enzyme conjugates according to the specific requirements in a variety of applications including biosensors, bioimaging, and biomedicine.

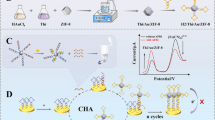
A self-assembled method has been presented for the facile and green synthesis of SA-β-Gal-CaHPO4 nanocomplexes with flower-like shape and high activity, and further employed as signal tag for fluorescent sensing of protein biomarker.






Similar content being viewed by others
References
Somturk B, Hancer M, Ocsoy I, Ozdemir N. Synthesis of copper ion incorporated horseradish peroxidase-based hybrid nanoflowers for enhanced catalytic activity and stability. Dalton Trans. 2015;44:13845–52.
Mittal S, Kaur H, Gautam N, Mantha AK. Biosensors for breast cancer diagnosis: a review of bioreceptors, biotransducers and signal amplification strategies. Biosens Bioelectron. 2017;88:217–31.
Zhou Z, Hartmann M. Progress in enzyme immobilization in ordered mesoporous materials and related applications. Chem Soc Rev. 2013;42:3894–912.
Ming L, Youzeng F, Tingwei W, Jianguo G. Micro-/nanorobots at work in active drug delivery. Adv Funct Mater. 2018. https://doi.org/10.1002/adfm.201706100.
Spicer CD, Jumeaux C, Gupta B, Stevens MM. Peptide and protein nanoparticle conjugates: versatile platforms for biomedical applications. Chem Soc Rev. 2018;47:3574–620.
Fang Y, Xu W, Wu J, Xu Z-K. Enzymatic transglycosylation of PEG brushes by β-galactosidase. Chem Commun. 2012;48:11208–10.
Tang F, Li L, Chen D. Mesoporous silica nanoparticles: synthesis, biocompatibility and drug delivery. Adv Mater. 2012;24:1504–34.
Jia H. Enzyme-carrying electrospun nanofibers. In: Wang P, editor. Nanoscale biocatalysis: methods and protocols. Totowa: Humana Press; 2011. p. 205–12.
Kalska-Szostko B, Rogowska M, Satuła D. Organophosphorous functionalization of magnetite nanoparticles. Colloids Surf B. 2013;111:656–62.
Wang Q, Zhou L, Jiang Y, Gao J. Improved stability of the carbon nanotubes–enzyme bioconjugates by biomimetic silicification. Enzym Microb Technol. 2011;49:11–6.
Ge J, Lei J, Zare RN. Protein-inorganic hybrid nanoflowers. Nat Nanotechnol. 2012;7:428–32.
Wang X, Shi J, Li Z, Zhang S, Wu H, Jiang Z. Facile one-pot preparation of chitosan/calcium pyrophosphate hybrid microflowers. ACS Appl Mater Interfaces. 2014;6:14522–32.
Hua X, Xing Y, Zhang X. Enhanced promiscuity of lipase-inorganic nanocrystal composites in the epoxidation of fatty acids in organic media. ACS Appl Mater Interfaces. 2016;8:16257–61.
Thawari AG, Rao CP. Peroxidase-like catalytic activity of copper-mediated protein–inorganic hybrid nanoflowers and nanofibers of β-lactoglobulin and α-lactalbumin: synthesis, spectral characterization, microscopic features, and catalytic activity. ACS Appl Mater Interfaces. 2016;8:10392–402.
Cui J, Zhao Y, Liu R, Zhong C, Jia S. Surfactant-activated lipase hybrid nanoflowers with enhanced enzymatic performance. Sci Rep. 2016;6:27928.
Li M, Luo M, Li F, Wang W, Liu K, Liu Q, et al. Biomimetic copper-based inorganic–protein nanoflower assembly constructed on the nanoscale fibrous membrane with enhanced stability and durability. J Phys Chem C. 2016;120:17348–56.
Zhu X, Huang J, Liu J, Zhang H, Jiang J, Yu R. A dual enzyme-inorganic hybrid nanoflower incorporated microfluidic paper-based analytic device (μPAD) biosensor for sensitive visualized detection of glucose. Nanoscale. 2017;9:5658–63.
Liu Y, Chen J, Du M, Wang X, Ji X, He Z. The preparation of dual-functional hybrid nanoflower and its application in the ultrasensitive detection of disease-related biomarker. Biosens Bioelectron. 2017;92:68–73.
Ariza-Avidad M, Salinas-Castillo A, Capitán-Vallvey LF. A 3D μPAD based on a multi-enzyme organic–inorganic hybrid nanoflower reactor. Biosens Bioelectron. 2016;77:51–5.
Li W, Lu S, Bao S, Shi Z, Lu Z, Li C, et al. Efficient in situ growth of enzyme-inorganic hybrids on paper strips for the visual detection of glucose. Biosens Bioelectron. 2018;99:603–11.
Sun J, Ge J, Liu W, Lan M, Zhang H, Wang P, et al. Multi-enzyme co-embedded organic-inorganic hybrid nanoflowers: synthesis and application as a colorimetric sensor. Nanoscale. 2014;6:255–62.
Guan Z, Zou Y, Zhang M, Lv J, Shen H, Yang P, et al. A highly parallel microfluidic droplet method enabling single-molecule counting for digital enzyme detection. Biomicrofluidics. 2014;8(1):014110.
Martinez-Bilbao M, Gaunt MT, Huber RE. E461H-beta-galactosidase (Escherichia coli): altered divalent metal specificity and slow but reversible metal inactivation. Biochemistry. 1995;34:13437–42.
Funding
This work was supported by the National Natural Science Foundation of China (21475101, 21675119).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical standards and informed consent
The study was approved by the Ethical Committee of Wuhan University. Human fluid samples used in this study do not have any identifying information about all the participants that provided written informed consent.
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 917 kb)
Rights and permissions
About this article
Cite this article
Liu, Y., Wang, B., Ji, X. et al. Self-assembled protein-enzyme nanoflower-based fluorescent sensing for protein biomarker. Anal Bioanal Chem 410, 7591–7598 (2018). https://doi.org/10.1007/s00216-018-1398-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-018-1398-7