Abstract
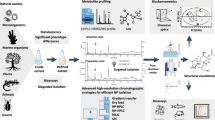
Development of fast and accurate methods to discover lead compounds for drug candidates is highly important. In this study, a reliable and effective post-column on-flow biochemical assay (POBA) was established to screen potent peroxidase inhibitors from complex chemical mixtures (e.g., natural product extracts). Multiple factors such as flow rate, organic phase, detection wavelength, and reaction coil were carefully investigated. To better understand the features of POBA, another emerging technology of ultrafiltration LC-MS was used for comparison. The result showed that POBA had advantages in saving time, avoiding false positives, and improving the accuracy. To illustrate the practicality of the method, Radix Salvia Miltiorrhizae, a traditional herb for cardiovascular disease treatment, was applied as the research objective. Finally, six compounds including tanshinol, protocatechuic aldehyde, salvianolic acid D, rosmarinic acid, lithospermic acid, and salvianolic acid B were determined as novel peroxidase inhibitors. Their bioactivities were validated by microplate-based assay, molecular docking, and pharmacophore modeling. This study demonstrates a great potential of POBA in the efficient and accurate discovery of drug candidates.

Compared with a classical method of ultrafiltration LC-MS, the newly developed method of on-flow bioassay shows advantages in saving time, avoiding false positives and improving the accuracy






Similar content being viewed by others
References
Song HP, Chen J, Hong JY, Hao HP, Qi LW, Lu J, et al. A strategy for screening of high-quality enzyme inhibitors from herbal medicines based on ultrafiltration LC-MS and in silico molecular docking. Chem Commun. 2015;51:1494–7.
Yang ZZ, Zhang YF, Sun LJ, Wang Y, Gao XM, Cheng YY. An ultrafiltration high-performance liquid chromatography coupled with diode array detector and mass spectrometry approach for screening and characterizing tyrosinase inhibitors from mulberry leaves. Anal Chim Acta. 2012;719:87–95.
Choi Y, Jermihov K, Nam SJ, Sturdy M, Maloney K, Qiu X, et al. Screening natural products for inhibitors of quinone reductase-2 using ultrafiltration LC-MS. Anal Chem. 2011;83:1048–52.
Tao Y, Cai H, Li WD, Cai BC. Ultrafiltration coupled with high-performance liquid chromatography and quadrupole-time-of-flight mass spectrometry for screening lipase binders from different extracts of Dendrobium officinale. Anal Bioanal Chem. 2015;407:6081–93.
Zheng C, Zhang XL, Liu W, Liu B, Yang HH, Lin ZA, et al. A selective artificial enzyme inhibitor based on nanoparticle-enzyme interactions and molecular imprinting. Adv Mater. 2013;25:5922–7.
Cutivet A, Schembri C, Kovensky J, Haupt K. Molecularly imprinted microgels as enzyme inhibitors. J Am Chem Soc. 2009;141:14699–702.
de Almeida FG, Vanzolini KL, Cass QB. Angiotensin converting enzyme immobilized on magnetic beads as a tool for ligand fishing. J Pharm Biomed Anal. 2017;132:159–64.
Cao J, Xu JJ, Liu XG, Wang SL, Peng LQ. Screening of thrombin inhibitors from phenolic acids using enzyme-immobilized magnetic beads through direct covalent binding by ultrahigh-performance liquid chromatography coupled with quadrupole time-of-flight tandem mass spectrometry. J Chromatogr A. 2016;1468:86–94.
van Breemen RB, Tao Y, Li WK. Cyclooxygenase-2 inhibitors in ginger (Zingiber officinale). Fitoterapia. 2011;82:38–43.
Wang HB, Zhao XP, Wang SF, Tao S, Ai N, Wang Y. Fabrication of enzyme-immobilized halloysite nanotubes for affinity enrichment of lipase inhibitors from complex mixtures. J Chromatogr A. 2015;1392:20–7.
Song HP, Zhang H, Fu Y, Mo HY, Zhang M, Chen J, et al. Screening for selective inhibitors of xanthine oxidase from Flos Chrysanthemum using ultrafiltration LC-MS combined with enzyme channel blocking. J Chromatogr B. 2014;961:56–61.
Wei H, Zhang XJ, Tian X, Wu GH. Pharmaceutical applications of affinity-ultrafiltration mass spectrometry: recent advances and future prospects. J Pharm Biomed Anal. 2016;131:444–53.
Potterat O, Hamburger M. Concepts and technologies for tracking bioactive compounds in natural product extracts: generation of libraries, and hyphenation of analytical processes with bioassays. Nat Prod Rep. 2013;30:546–64.
Newman DJ, Cragg GM. Natural products as sources of new drugs from 1981 to 2014. J Nat Prod. 2016;79:629–61.
Rishton GM. Natural products as a robust source of new drugs and drug leads: past successes and present day issues. Am J Cardiol. 2008;101:43–9.
Deng Y, Ding CY, Ye N, Liu ZQ, Wold EA, Chen HY, et al. Discovery and development of natural product oridonin-inspired anticancer agents. Eur J Med Chem. 2016;122:102–17.
Mishra BB, Tiwari VK. Natural products: an evolving role in future drug discovery. Eur J Med Chem. 2011;46:4769–807.
Harvey AL, Edrada-Ebel R, Quinn RJ. The re-emergence of natural products for drug discovery in the genomics era. Nat Rev Drug Discov. 2015;14:111–29.
Regasini LO, Vellosa JC, Silva DH, Furlan M, de Oliveira OMM, Khalil NM, et al. Flavonols from Pterogyne nitens and their evaluation as myeloperoxidase inhibitors. Phytochemistry. 2008;69:1739–44.
Eiserich JP, Baldus S, Brennan ML, Ma W, Zhang C, Tousson A, et al. Myeloperoxidase, a leukocyte-derived vascular NO oxidase. Science. 2002;296:2391–4.
Fu XY, Kassim SY, Parks WC, Heinecke JW. Hypochlorous acid oxygenates the cysteine switch domain of pro-matrilysin (MMP-7). A mechanism for matrix metalloproteinase activation and atherosclerotic plaque rupture by myeloperoxidase. J Biol Chem. 2001;276:41279–87.
Malle E, Furtmuller PG, Sattler W, Obinger C. Myeloperoxidase: a target for new drug development? Br J Pharmacol. 2007;152:838–54.
Wang JG, Mahmud SA, Thompson JA, Geng JG, Key NS, Slungaard A. The principal eosinophil peroxidase product, HOSCN, is a uniquely potent phagocyte oxidant inducer of endothelial cell tissue factor activity: a potential mechanism for thrombosis in eosinophilic inflammatory states. Blood. 2006;107:558–65.
Cheng HT, Li XL, Li XR, Li YH, Wang LJ, Xue M. Simultaneous quantification of selected compounds from Salvia herbs by HPLC method and their application. Food Chem. 2012;130:1031–5.
Doerge DR, Divi RL, Churchwell MI. Identification of the colored guaiacol oxidation product produced by peroxidases. Anal Biochem. 1997;250:10–7.
Lavery CB, Macinnis MC, Macdonald MJ, Williams JB, Spencer CA, Burke AA, et al. Purification of peroxidase from horseradish (Armoracia rusticana) roots. J Agric Food Chem. 2010;58:8471–6.
Yeoh BS, Aguilera Olvera R, Singh V, Xiao X, Kennett MJ, Joe B, et al. Epigallocatechin-3-gallate inhibition of myeloperoxidase and its counter-regulation by dietary iron and lipocalin 2 in murine model of gut inflammation. Am J Pathol. 2016;186:912–26.
Song HP, Wu SQ, Hao HP, Chen J, Lu J, Xu XJ, et al. A chemical family-based strategy for uncovering hidden bioactive molecules and multicomponent interactions in herbal medicines. Sci Rep. 2016;6:23840–54.
Przyjazny A, Kjellstrom TL, Bachas LG. High-performance liquid chromatographic postcolumn reaction detection based on a competitive binding system. Anal Chem. 1990;62:2536–40.
de Boer AR, Letzel T, van Elswijk DA, Lingeman H, Niessen WM, Irth H. On-flow coupling of high-performance liquid chromatography to a continuous-flow enzyme assay based on electrospray ionization mass spectrometry. Anal Chem. 2004;76:3155–61.
Kool J, de Kloe GE, Bruyneel B, de Vlieger JS, Retra K, Wijtmans M, et al. Online fluorescence enhancement assay for the acetylcholine binding protein with parallel mass spectrometric identification. J Med Chem. 2010;53:4720–30.
Li DQ, Zhao J, Li SP. High-performance liquid chromatography coupled with post-column dual-bioactivity assay for simultaneous screening of xanthine oxidase inhibitors and free radical scavengers from complex mixture. J Chromatogr A. 2014;1345:50–6.
Kool J, van Liempd SM, Ramautar R, Schenk T, Meerman JH, Irth H, et al. Development of a novel cytochrome p450 bioaffinity detection system coupled online to gradient reversed-phase high-performance liquid chromatography. J Biomol Screen. 2005;10:427–36.
Han JY, Fan JY, Horie Y, Miura S, Cui DH, Ishii H, et al. Ameliorating effects of compounds derived from Salvia miltiorrhiza root extract on microcirculatory disturbance and target organ injury by ischemia and reperfusion. Pharmacol Ther. 2008;117:280–95.
Guo Y, Li Y, Xue L, Severino RP, Gao S, Niu J, et al. Salvia miltiorrhiza: an ancient Chinese herbal medicine as a source for anti-osteoporotic drugs. J Ethnopharmacol. 2014;155:1401–16.
Chen XF, Lou ZY, Zhang H, Tan GG, Liu ZR, Li WH, et al. Identification of multiple components in Guanxinning injection using hydrophilic interaction liquid chromatography/time-of-flight mass spectrometry and reversed-phase liquid chromatography/time-of-flight mass spectrometry. Rapid Commun Mass Spectrom. 2011;25:1661–74.
Zhu JF, Yi XJ, Zhang JH, Chen SQ, Wu YJ. Chemical profiling and antioxidant evaluation of Yangxinshi tablet by HPLC-ESI-Q-TOF-MS/MS combined with DPPH assay. J Chromatogr B. 2017;1060:262–71.
Zhang H, Wang SQ, Liu Y, Luo LP, Liu P, Qi LW, et al. Trace analysis in complex mixtures using a high-component filtering strategy with liquid chromatography-mass spectrometry. J Pharm Biomed Anal. 2012;70:169–77.
Howes BD, Heering HA, Roberts TO, Schneider-Belhadadd F, Smith AT, Smulevich G. Mutation of residues critical for benzohydroxamic acid binding to horseradish peroxidase isoenzyme C. Biopolymers. 2001;62:261–7.
Acknowledgements
This work was supported by the "Twelfth-Five years" Supporting Programs from the Ministry of Science and Technology of China (No. 2015ZX09101043-010), the Foundation for Innovative Research Groups of the National Natural Science Foundation of China (Grant No. 81421005) and a Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 1844 kb)
Rights and permissions
About this article
Cite this article
Wu, SQ., Song, HP., Li, B. et al. A fast and accurate method for the identification of peroxidase inhibitors from Radix Salvia Miltiorrhizae by on-flow biochemical assay coupled with LC/Q-TOF-MS: comparison with ultrafiltration-based affinity selection. Anal Bioanal Chem 410, 4311–4322 (2018). https://doi.org/10.1007/s00216-018-1081-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-018-1081-z