Abstract
The hierarchy of human hemopoietic progenitor cells that produce lymphoid and granulocytic–monocytic (myeloid) lineages is unclear. Multiple progenitor populations produce lymphoid and myeloid cells, but they remain incompletely characterized. Here we demonstrated that lympho-myeloid progenitor populations in cord blood — lymphoid-primed multi-potential progenitors (LMPPs), granulocyte-macrophage progenitors (GMPs) and multi-lymphoid progenitors (MLPs) — were functionally and transcriptionally distinct and heterogeneous at the clonal level, with progenitors of many different functional potentials present. Although most progenitors had the potential to develop into only one mature cell type (‘uni-lineage potential’), bi- and rarer multi-lineage progenitors were present among LMPPs, GMPs and MLPs. Those findings, coupled with single-cell expression analyses, suggest that a continuum of progenitors execute lymphoid and myeloid differentiation, rather than only uni-lineage progenitors’ being present downstream of stem cells.
Similar content being viewed by others
Main
Human hemopoiesis produces 10 × 109 new, terminally mature blood cells daily, production that is also rapidly responsive to external change. Most of this production generates red cells, short-lived myeloid cells and platelets. It also replenishes long-lived acquired immune cells and natural killer (NK) cells of the innate immune system. Dysregulation of this complex process can lead to hemopoietic deficiency, immunodeficiency and blood cancer. Active debate continues about the heterogeneity and plasticity of hemopoietic cell populations at steady state and in response to stimuli. At the hierarchy apex lie multi-potent hemopoietic stem cell (HSC) populations, which are heterogeneous in their differentiation potential, cell cycle, self-renewal capacity, stability over time and contribution to hemopoiesis at steady state versus during transplantation1,2,3,4,5,6,7,8,9,10,11. Downstream of long-term HSCs in the mouse are heterogeneous short-term HSCs, multipotent progenitors (MPPs) and early lineage-biased progenitors5,7,12,13,14. The short-term HSC–MPP populations in humans have not been fully defined15,16. In terms of lineage-potential restriction, the erythroid and megakaryocyte fates probably diverge early from other myeloid and lymphoid potentials in mice14,17,18,19,20 and humans21,22,23,24,25 and might arise directly from either HSCs6 or downstream MPPs14,16,26.
Focusing on the first human lympho-myeloid progenitors downstream of HSCs and MPPs, two progenitor populations have been identified in the immature Lin–CD34+CD38–CD90neg–lo compartment. These include Lin–CD34+CD38–CD90neg–loCD45RA+CD10– lymphoid-primed multi-potential progenitors (LMPPs) with granulocytic, monocytic, B cell and T cell potential but that are unable to generate erythrocytes or megakaryocytes22. Such data support published studies showing that human CD34+CD10– cells retain lympho-myeloid potential and progressively lose myeloid potential with expression of the cell-surface marker CD10 (refs. 27,28). In contrast, the multi-lymphoid progenitors (MLPs), initially reported as Lin–CD34+CD38–CD90neg–loCD45RA+CD10+ cells, have lymphoid (B cell, T cell and NK cell), monocytic and dendritic cell (DC) potential but cannot make granulocytes21. However, CD10– MLP populations29 have been reported that might overlap the LMPPs. Within the Lin–CD34+CD38+CD45RA+ compartment, there are at least two lympho-myeloid progenitors: a CD62LhiCD10– lymphoid-primed progenitor with lymphoid, monocytic and DC potential23, and the granulocyte-monocyte progenitor (GMP) (Lin–CD34+CD38+CD45RA+CD123+). The GMP population includes both a CD62Lhi subpopulation and a CD62Llo subpopulation and has mainly myeloid potential but retains residual lymphoid potential22,30, consistent with the mouse pre-granulocyte-monocyte progenitor31. Finally, the human Lin–CD34+CD38+CD45RA+ compartment also includes a CD10+ subpopulation with T cell, B cell, NK cell and DC potential but lacking myeloid potential32. Such observations have raised questions about whether these progenitor populations are pure or heterogeneous, how distinct they are and the nature of the functional, transcriptional and hierarchical relationships among them.
Lympho-myeloid progenitors in the Lin–CD34+ CD45RA+ CD90neg–lo compartment that can be either CD38+ or CD38– and either CD10+ or CD10– have been described. Here we directly and rigorously compared the in vitro and in vivo functional potential and transcriptional programs of human LMPPs, MLPs and GMPs. We found that these progenitors were distinct and heterogeneous. Single-cell gene-expression analysis indicated a continuum of progenitors with lymphoid and myeloid potential downstream of HSCs. The bulk of the multi-lineage lympho-myeloid progenitors were in a sub-compartment of Lin–CD34+CD38–CD90neg–loCD45RA+CD10– LMPPs (called ‘LMPPmix cells’ here) with the lowest expression, among LMPPs, of the cell-surface markers CD10 and CD45RA.
Results
In vitro assays of the potential of distinct lympho-myeloid progenitors
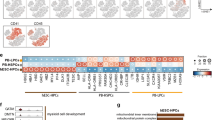
We improved published flow-cytometry staining and sorting strategies21,22 to prospectively purify eight human hemopoietic stem–progenitor cell (HSPC) populations (Supplementary Table 1 and Supplementary Fig. 1a) in human cord blood (CB) and bone marrow (BM). These HSPC populations included the following: HSCs (Lin–CD34+CD38–CD90+CD45RA–CD10–), MPPs (Lin–CD34+CD38–CD90–CD45RA–CD10–), LMPPs (Lin–CD34+CD38–CD90neg–loCD45RA+CD10–), MLPs (Lin–CD34+CD38–CD90neg–loCD45RA+CD10+), common myeloid progenitors (CMPs) (Lin–CD34+CD38+CD123+CD45RA–CD10–), GMPs (Lin–CD34+CD38+CD123+CD45RA+CD10–), megakaryocyte erythroid progenitor (MEPs) (Lin–CD34+CD38+CD123–CD45RA–CD10–), and B cell–NK cell (B-NK) progenitors (Lin–CD34+CD38+CD90–CD45RA+CD10+) (Supplementary Table 1 and Supplementary Fig. 1a). Among CB Lin–CD34+ cells, these eight HSPC populations accounted for 82% of cells; the remaining cells did not constitute separate populations (Supplementary Table 1 and Supplementary Fig. 1a). The Lin–CD34+CD38–CD45RA+ compartment contained a mixture of CD10– LMPPs and CD10+ MLPs (Supplementary Fig. 1a). Furthermore, the more-mature Lin–CD34+CD38+ compartment was separated into the CD10+ B-NK progenitor population32 and CD10– heterogeneous myeloid progenitors (Supplementary Fig. 1a). Immunophenotypic LMPPs and MLPs were rare (Supplementary Table 1). As assessed with analysis gates, they constituted 0.2% of the BM Lin–CD34+ compartment and ~2 cells among 1 × 105 BM mononuclear cells (MNCs) (Supplementary Table 1). Although they were more frequent in CB, they still constituted only ~1 cell among 1 × 104 CB MNCs (Supplementary Table 1). GMPs were 20-fold more abundant in CB and 100-fold more abundant in BM than were LMPPs or MLPs (~1.5–2 cells among 1 × 103 MNCs) (Supplementary Table 1).
As the frequency of adult BM LMPPs and MLPs was extremely low, we used fresh CB cells as a source of HSPCs. Cells were double-sorted to high purity (of >99%, except CMPs, with a purity of 97%). In methylcellulose-based colony-forming-unit assays, the LMPPs and MLPs had lower myeloid clonogenic potential (6% (LMPPs) and <1% (MLPs)) than that of GMPs (31%) (Fig. 1a,b). GMPs and LMPPs generated granulocyte colonies, monocyte-macrophage colonies and granulocyte and monocyte-macrophage colonies with no or minimal erythrocyte potential (<0.5%) (Fig. 1a). MLPs only generated very few monocyte colonies (Fig. 1a), consistent with published data15,21,22,33.
a, Colony-forming unit (CFU) assay of the cloning efficiency and myelo-erythroid lineage affiliation of CB HSPCs (150 cells plated; horizontal axis), showing the frequency of mixed erythro-myeloid colonies (CFU-mix), monocyte-macrophage colonies (CFU-M), granulocyte colonies (CFU-G), granulocyte and monocyte-macrophage colonies (CFU-GM) and erythroid colonies (burst-forming units–erythroid and CFU-E (E)) (key). b, Microscopy showing the morphology of cells from the CFU assay in a (left margin), stained with May–Grunwald–Giemsa (left), and flow cytometry of cells harvested from the colony types in a (middle and right). Scale bars (left), 10 µm. Numbers in outlined areas (middle and right) indicate percent CD15+CD14– neutrophil granulocytes (top left), CD15–CD14+ monocyte-macrophages (bottom right) or CD15–CD14– cells (bottom left) (middle column), or CD15–CD14–CD235+ erythroid cells (bottom right) (right column). c, Lineage output (key) of LMPPs, MLPs and GMPs (above plot; 150 each) after 1, 2 or 3 weeks (horizontal axis) in SGF15/2 culture, presented as frequency among hCD45+ cells (flow cytometry of 2-week cultures, Supplementary Fig. 1b). d, Gene expression by output cells from c (horizontal axis), purified by flow cytometry; results are presented relative to those of the control genes GAPDH and ACTB. e, T cell output by LMPPs, MLPs and GMPs (above plots) in bulk after 5 or 7 weeks (horizontal axis) in SF7a culture, showing the frequency of CD7+CD1a+ cells of the T cell lineage (left), and of CD7+CD1a+ CD4–CD8+ cells (CD8), CD7+CD1a+ CD4+CD8– cells (CD4), CD7+CD1a+ CD4+CD8+ cells (DP) and CD7+CD1a+ CD4–CD8– cells (DN) (key) (right) among hCD45+ cells (flow cytometry of 5-week cultures, Supplementary Fig. 1c). f, Gene expression by output T cells from e, purified by flow cytometry (left half of horizontal axis; as in key in e), and by control mature non-T cells obtained from the sorting of cells from E, G and M colonies from a (right half of horizontal axis); results are presented relative to those of GAPDH and ACTB. Data are representative of three experiments (a; error bars, mean + s.d. of five donors), two experiments (b), two experiments (c; mean + s.d of three donors), two experiments (d; mean + s.e.m. of three donors), three experiments (e; mean + s.d. of five donors) or three experiments (f; mean + s.e.m. of three donors).
We analyzed the lymphoid and myeloiddifferentiation potential of a population of 150 LMPPs, MLPs and GMPs using an optimized, new in vitro liquid culture on MS-5 mouse stromal cells supplemented with the cytokines SCF, G-CSF, FLT3L, IL-2 and IL-15, and the prostaglandin-synthesis inhibitor DUP-697 (called ‘SGF15/2 culture’ here). By performing a kinetic analysis of lineage output, we determined that 2 weeks was the optimal time at which to detect human CD45+CD15+ (hCD45+CD15+) neutrophil-granulocytes (Gs), hCD45+CD14+ monocyte-macrophages (Ms), hCD45+CD19+ B cells and hCD45+CD56+ NK cells in the culture (Fig. 1c and Supplementary Fig. 1b). Gs, Ms, B cells or NK cells purified by flow cytometry from the in vitro SGF15/2 culture expressed the appropriate lineage-associated genes (Fig. 1d and Supplementary Table 2). Therefore, we analyzed all subsequent limiting-dilution and single-cell cultures at week 2 to capture all four outputs: myeloid (G and M) and lymphoid (B and NK). We assessed the production of T cells by LMPP, MLP and GMP populations at weeks 5 and 7 using an in vitro liquid culture assay on OP9 stromal cells expressing the human Notch receptor ligand DL1 (OP9-hDL1 cells) in the presence of SCF, FLT3L and IL-7 (called ‘SF7a culture’ here) (Fig. 1e and Supplementary Fig. 1c). LMPPs, GMPs and MLPs generated hCD7+CD1a+ cells of the T cell lineage, including hCD7+CD1a+ CD4+CD8+ (double-positive) T cells and hCD7+CD1a+ CD4–CD8+ or CD4+CD8– (single-positive) T cells (Fig. 1e and Supplementary Fig. 1c). T cell subpopulations purified by flow cytometry expressed the appropriate lineage-associated genes (Fig. 1f and Supplementary Table 2). In summary, we established conditions with which to prospectively purify eight HSPC populations and to test their in vitro potential to develop into myeloid and lymphoid lineages.
Functional heterogeneity of lympho-myeloid progenitors
We next used four in vitro liquid culture assays to assess the clonal potential of CB LMPPs, MLPs and GMPs (Supplementary Fig. 2a,b). First, we performed limiting-dilution assays of SGF15/2 cultures and assessed lineage output by flow cytometry (Supplementary Fig. 2c). One in two LMPPs produced B cells and one in three LMPPs produced NK cells, but only one in five LMPPs produced monocytes and one in ten LMPPs produced granulocytes (Table 1 and Supplementary Fig. 2d). GMPs generated myeloid cells with higher frequency than did LMPPs (one in two for Ms and one in four for Gs) and generated lymphoid cells at lower frequency than did LMPPs (one in twenty-two for B cells and one in eight for NK cells) (Table 1 and Supplementary Fig. 2d). 1 in 11 MLP cells produced B cells and 1 in 18 generated NK cells, whereas myeloid output was rare (Table 1), which indicated that MLPs were lymphoid biased. Bi-lineage and multi-lineage cells were detected at a lower frequency (1 in 6 to 1 in 789) (Table 1 and Supplementary Table 3a).
As the limiting-dilution assay does not rigorously define the frequency of multi-lineage functional potential at a clonal level, we assessed lympho-myeloid (B, NK, G and M) potential in the optimized SGF15/2 culture. We assessed the potential of 1,136 LMPPs, 710 MLPs and 1,622 GMPs as single cells, isolated from 22 CB donors (a total of 6.3 × 109 MNCs), to provide robust quantitative data, especially for rare functional potentials (Supplementary Table 3b). At the single-cell level, LMPPs and GMPs had higher cloning efficiency (54% and 71%, respectively) than that of MLPs (11%) (Fig. 2a). LMPPs and MLPs were mainly lymphoid progenitors, whereas GMPs were mainly myeloid progenitors (Fig. 2a). Focusing on productive wells (defined as wells that produced any type of mature output), we found that 69% of LMPPs, 88% of MLPs and 63% of GMPs produced uni-lineage output (Fig. 2b). When there was uni-lineage output, 92% of LMPPs had lymphoid output (B or NK) and 8% had myeloid output (G or M) (Fig. 2b). The MLPs were almost exclusively lymphoid progenitors with very low myeloid output (3%) (Fig. 2b). 79% of GMP cells had myeloid output, and 21% had lymphoid output (Fig. 2b). Bi-lineage output was detected in 24% of LMPPs, 12% of MLPs and 33% of GMPs (Fig. 2c), and output of three or more lineages was rare (6% of LMPPs, 0% of MLPs and 3% of GMPs) (Fig. 2d). Only 8% of all plated LMPPs, 7% of GMPs and hardly any MLPs (0.3%) exhibited combined lympho-myeloid potential (Fig. 2e). The lympho-myeloid output of LMPPs was significantly higher than that of GMPs (P = 0.0125) or MLPs (P = 0.0019, Supplementary Table 3c).
a, Total cloning efficiency of single LMPPs (615 of 1,136 cells), MLPs (76 of 710 cells) and GMPs (1,145 of 1,622 cells) in SGF15/2 culture (left), and cloning efficiency of lymphoid lineages (middle) and myeloid lineages (right), showing total cloning efficiency (Total) and proportion of lymphoid potential (lymphoid plus mixed; Ly) or myeloid potential (myeloid plus mixed clones; My) (key). b–d, Uni-lineage output (b), bi-lineage output (c) and multi-lineage output (d) of single CB LMPPs, MLPs and GMPs (horizontal axis) in SGF15/2 culture, presented as the proportion of cells of various lineages (key) in wells that produced cells (positive wells). e, Lymphoid, myeloid and lympho-myeloid (Ly-My) output (key) of LMPPs, MLPs and GMPs (horizontal axis) in SGF15/2 culture, presented as the frequency among all plated cells. f, Total cloning efficiency of single CB LMPPs (128 of 215 cells), MLPs (37 of 197 cells) and GMPs (127 of 219 cells) in SF7b/Dox culture (left), and cloning efficiency of lymphoid lineages (middle) and myeloid lineages (right), showing total cloning efficiency and proportion of lymphoid or myeloid potential (key). g–i, Uni-lineage output (g), bi-lineage output (h) and multi-lineage output (i) of single CB LMPPs, MLPs and GMPs (horizontal axis) in SF7b/Dox culture, presented as the frequency of cells of various lineages (key) in positive wells. j, Lymphoid, myeloid and lympho-myeloid output (key) of LMPPs, MLPs and GMPs (horizontal axis) in SF7b/Dox culture, presented as the frequency among all plated cells. NS, not significant (P > 0.05); *P = 0.01–0.05, **P = 0.001–0.01 and ***P < 0.001 (Fisher’s exact test (a,f, left), Students t-test (a,f, middle and right). Data are representative of nine experiments with 22 donors (a, left, and b–e), nine experiments (a, middle and right; mean + s.d. of 22 donors), one experiment with three donors (f, left, and g–j) or one experiment (f, middle and right; mean + s.d. of three donors).
We also assessed the lympho-myeloid (B, NK, G and M) potential of 96 LMPPs, 52 MLPs and 110 GMPs as single cells in a third in vitro liquid culture assay with MS-5 stroma in the presence of the cytokines SCF, IL-7, thrombopoietin, IL-2, GM-CSF, G-CSF and M-CSF (called ‘S7T2GM/G/M culture’ here) that was used to define the MLPs21 (Supplementary Fig. 2b). We obtained results similar to those of SGF15/2 culture, but S7T2GM/G/M cultures were less permissive for granulocytic output (Supplementary Fig. 2e–i and Supplementary Table 3d). Most of the output from LMPPs was uni-lineage, with rarer bi-lineage and less frequent multi-lineage output (Supplementary Fig. 2e–i and Supplementary Table 3d). MLPs exhibited only lymphoid uni-lineage output (Supplementary Fig. 2e–i and Supplementary Table 3d).
Finally, we assessed the lympho-myeloid potential of 215 LMPPs, 197 MLPs and 219 GMPs as single cells in a fourth assay, an independent culture condition optimized for the detection of combined lymphoid (B, NK and T) and myeloid (M and G) potential. Single LMPPs, MLPs and GMPs were cultured in the presence of SCF, FLT3L and IL-7 on an MS-5 stromal cell line that can be induced to express hDL1 by doxycycline treatment (MS-5–hDL1IND stroma) (called ‘SF7b/Dox culture’ here). The B-NK-M-G output was analyzed at 3 weeks and the T cell output was analyzed at 6 weeks (Fig. 2f–j and Supplementary Table 3e). Uni-lineage T cell output was detected in LMPP and MLP populations (3% of productive wells) but was almost completely absent in GMP populations (<0.1%) (Fig. 2g). T cell output combined with other lymphoid output was detected for 1–5% of LMPPs and MLPs and rarely for GMPs (Fig. 2g). Lympho-myeloid output was detected only for LMPPs (14%) (Fig. 2i). Overall, 24 functionally different progenitor types were identified in the three single-cell in vitro clonal assays; all 24 progenitor types were observed for LMPPs, and only subsets of those were noted for MLPs and GMPs (Supplementary Fig. 2j).
Ossicle assay defines the in vivo potential of LMPPs, MLPs and GMPs
Successful single-cell transplantation of human progenitors in xenotransplantation assays is not feasible. Furthermore, direct injection of progenitor populations into immunodeficient mice yields low engraftment (< 0.1%)15,21,22,25. Therefore, we assessed in vivo progenitor function in a new humanized ossicle model34. Human BM-derived mesenchymal stromal cells were subcutaneously injected into immunodeficient mice, where they form a humanized ossicle over 8 weeks. LMPPs, MLPs and GMPs were injected into the ossicle, and lineage output was analyzed 1 and 2 weeks after injection (Supplementary Fig. 3a,b). Engraftment was detected at both time points, with greater engraftment of hCD45+hCD33+hCD14+ cells (M), hCD45+hCD33+hCD15+ cells (G) and hCD45+hCD33–hCD19+ cells (B) at week 2 than at week 1 (data not shown). All subsequent analyses were done at 2 weeks after transplantation. As the number of cells injected varied (~300–60,000 cells, depending on the progenitor subset; Supplementary Fig. 3c), we report the mean cell engraftment per 1,000 transplanted cells here. GMPs had the greatest mean engraftment (2.6%), followed by LMPPs (1.4%) and MLPs (0.2%) (Fig. 3a). LMPPs produced more CD33+ myeloid cells (82%) than CD19+ B cells (17%) (Fig. 3b,c). MLPs generated more B cells (78% ± 5.9%) than myeloid cells (19% ± 6.7%) (mean ± s.e.m.) (Fig. 3b,c). There was no correlation between the number of the transplanted cells and the ratio of lymphoid cells to myeloid cells (Supplementary Fig. 3d). GMPs generated mainly myeloid cells (97%) (Fig. 3b,c). Myeloid cells generated from LMPPs and GMPs expressed markers of monocytes (CD14) and granulocytes (CD15) (Fig. 3c). No CD14+ or CD15+ cells were detected as being produced from MLPs (Fig. 3c). Morphological analysis of the engrafted cells confirmed that CD15+ cells were granulocytic and CD14+ cells were monocytic (Fig. 3d). CD14+CD15+ cells, generated by LMPPs and GMPs (Fig. 3c), were immature myeloid cells by morphology (Fig. 3d). Thus, LMPPs, MLPs and GMPs had different lymphoid potential and myeloid potential in the humanized ossicle assay.
a, Engraftment of human LMPPs, MLPs and GMPs (horizontal axis) into humanized ossicles in immunodeficient mice 2 weeks after transplantation, normalized to 1,000 transplanted cells and presented as percent human chimerism. b, Frequency of B cells and myeloid cells (key) among human CD45+HLA-ABC+ cells generated by LMPPs, MLPs and GMPs (horizonal axis) after transplantation as in a. c, Flow cytometry of cells from host mice at 2 weeks after transplantation as in a. Numbers in outlined areas indicate percent human CD45+HLA-ABC+ cells (human engraftment) (top row); CD19+ (B) cells (top left) or CD33+ (myeloid) cells (bottom right) (middle row); or CD15+CD14– granulocytes (top left), CD15+CD14+ immature myeloid cells (top right) or CD15–CD14+ monocytes (bottom right) (bottom row). d, Microscopy of CD15+CD14–, CD15+CD14+ and CD15–CD14+ myeloid cells (above images) generated by LMPPs at 2 weeks after transplantation as in a, stained with May–Grunwald–Giemsa. Scale bars, 20 μm. Each symbol (a, b) represents an individual recipient mouse; small horizontal lines indicate the mean. Data are from ten experiments (a–c; mean of n = 11 donors (LMPP), n = 3 donors (MLP) or n = 6 donors (GMP)) or two experiments with two donors (d).
LMPP, MLP and GMP transcriptional programs correlate with function
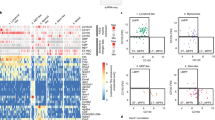
We performed RNA sequencing of human CB HSPC populations (HSC, MPP, LMPP, MLP, CMP, GMP and MEP). Hierarchical clustering using all expressed genes separated LMPPs and MLPs from the other HSPCs (Fig. 4a and Supplementary Fig. 4a). HSCs and MPPs clustered away from mature progenitors in this analysis (Fig. 4a and Supplementary Fig. 4a). We used analysis of variance (ANOVA) to identify genes with differential expression (‘differentially expressed genes’ (DEGs)) across the seven HSPC populations (Supplementary Table 4). We performed principal-component analysis (PCA) using all expressed genes or between 300 and 10,000 of the genes with the most significantly differential expression ('most significant DEGs'; ranked by ANOVA P value) (Fig. 4b). The best separation of HSPC populations on a PCA plot was achieved with the 300 most significant DEGs (Fig. 4b). Principal component 1 (PC1) separated HSPCs by lineage potential, and PC2 separated HSPCs by maturation (Fig. 4b). By comparing the eigenvalues of the 300 most significant DEGs against those from a randomized data set, we demonstrated that PC1–PC3 captured most of the variation between populations (Supplementary Fig. 4b,c). We also identified the genes with the greatest variance across all populations without assuming population identity (Fig. 4b and Supplementary Fig. 4d). PCA plotting using this gene set produced similar results (Fig. 4b and Supplementary Fig. 4d). The loadings plot for the PCA using the 300 most significant DEGs identified genes associated with stem cells (HLF, MECOM and NFIB), lymphoid cells (IGJ, IRF8 and MME) and erythroid-megakaryocytic cells (HBD and HPGDS) (Fig. 4c). Hierarchical clustering using that same gene set separated HSPC populations (Fig. 4d). GMPs had the highest expression of ELANE, MPO and PRTN3, whereas LMPPs and MLPs shared expression of many lymphoid-cell-associated genes (for example, IL-7R, LCK, SYK, ADA, HLX, LST1 and ITGAL) (Fig. 4d).
a, Hierarchical clustering of HSPC populations (key) using all genes and 1,000 bootstrap permutation analyses, with height calculated as (1 – [correlation co-efficient]); numbers above dendrogram bars in plot indicate approximate unbiased P values (au). b, PCA of CB HSPCs with all genes (top left) or the 300, 3,000 or 10,000 most significant DEGs, ranked by ANOVA P value (called, for example, ‘top 300 ANOVA genes’ subsequently; above plots), or the 300 genes with the greatest variance in expression (300 variant; bottom right); numbers in parentheses along axes indicate the proportion of variance represented by PC1 and PC2. c, Loadings plot showing the genes with the most extreme loadings scores for the PCA run with the top 300 ANOVA genes (left) or the 300 genes with the greatest variance in expression (right).d, Expression (key; normalized per gene) of the top 300 ANOVA genes (right margin; one per row) by HSPC populations (columns), showing clusters with distinct expression patterns across HSPC populations (yellow outlines); dendrograms (top and left margins) indicate hierarchical clustering (of columns and rows, respectively). e, Number of DEGs (key; specific number in plot) in various comparisons of HSPC populations (rows and columns). f,g, Expression (as in d) of the 50 most significant DEGs, ranked by P value, in the LMPP, MLP and GMP gene signatures (above plots) (f), and of transcription-factor-encoding genes expressed differentially across HSPC populations (g); asterisk colors indicate genes associated with the lymphoid lineage (orange) or myeloid lineage (green) or genes encoding products associated with immunological function (black). Data are representative of three experiments with three donors (MPPs) or four donors (all other cell populations).
Transcriptional relatedness between HSPC populations, without assumption of any hierarchical relationships, was further analyzed through pairwise comparisons. The most closely related populations were HSCs and MPPs (only 13 DEGs separated these populations), while LMPPs and MLPs were also closely related (85 DEGs) (Fig. 4e, Supplementary Fig. 4e and Supplementary Tables 5–11). GMPs were most closely related to CMPs (40 DEGs) but retained similarity to LMPPs (183 DEGs) (Fig. 4e, Supplementary Fig. 4e and Supplementary Tables 5–11). We derived gene-expression signatures for LMPPs, MLPs and GMPs from DEGs in one-versus-all population comparisons filtered for uniquely expressed genes (Fig. 4f and Supplementary Table 12a–c). The GMP signature included many myeloid-cell-associated genes, and the MLP signature included many lymphoid-cell-associated genes (Fig. 4d). In contrast, the LMPP signature included both lymphoid-cell-associated genes (ETS1, EBF1 and CYTIP) and myeloid-cell-associated genes (TRPM2, S100A8, PADI4 and ALOX15B) (Fig. 4f).
To confirm those findings, we investigated the profiles of LMPPs, GMPs and MLPs using published gene sets25. GMPs expressed genes associated with immature myeloid cells, whereas LMPPs and MLPs expressed genes associated with B cells, monocytes and DCs but not those associated with neutrophils (with the exception of FOSB) (Supplementary Fig. 4f). Additionally, in a MetaCore enrichment analysis, the GMPs showed enrichment for pathways (expressed a significant number of components of these pathways) associated with myeloid maturation (for example, granulocyte development (false-discovery rate (FDR) = 0.0136)), whereas MLPs showed enrichment for lymphopoiesis pathways (for example, Notch signaling (FDR < 0.001)) (Supplementary Table 13). LMPPs showed more balanced enrichment for both lymphoid pathways and myeloid pathways (for example, M-CSF signaling (FDR < 0.001) and BCR signaling (FDR = 0.049)) (Supplementary Table 13).
We used two approaches to pinpoint transcription factors driving the programs noted above. First, we identified transcription factors expressed differentially by MLPs relative to their expression by GMPs (Supplementary Fig. 4g). Second, we assessed the expression of hemopoietic transcription factors identified previously35 (Fig. 4g). In both analyses, GMPs expressed mainly genes encoding myeloid-cell-associated transcription factors (for example, ERG, GATA2, MYB and EGR1), while lymphoid-cell-associated transcription factors (for example, HES1, RUNX3, POU2F2, LEF1, IKZF1, IRF8, TCF4) had their highest expression in MLPs (Fig. 4g and Supplementary Fig. 4g). LMPPs showed expression of both lymphoid-cell-associated transcription factors and myeloid-cell-associated transcription factors (Fig. 4g and Supplementary Fig. 4g). A similar trend was seen for genes encoding cytokine and chemokine receptors (Supplementary Fig. 4h). Therefore, the transcriptional programs of LMPPs, MLPs and GMPs reflected their functional potential.
Single-cell RNA analysis reveals a continuum of differentiation
To begin to separate distinct progenitors in the heterogeneous GMP, LMPP and MLP populations, we sorted single cells by index flow cytometry for functional analysis, RNA sequencing and quantitative RT-PCR (qRT-PCR). Index data allowed correlation of function and transcriptional state36 (Fig. 5a). First, we profiled the expression of 96 genes encoding lineage-associated transcriptional regulators and cell-surface and lineage-associated markers (Supplementary Table 14) by a total of 919 single LMPPs, MLPs and GMPs. Genes with low variance and levels of detection were excluded. The expression of 74 genes was taken forward for analysis. Hierarchical clustering assigned GMPs, LMPPs and MLPs to three clusters: cluster 1 (543 cells) was mainly MLPs and LMPPs, cluster 2 (150 cells) was a mixture of GMPs, LMPPs and MLPs, and cluster 3 (226 cells) was mainly GMPs (Supplementary Fig. 5a,b). Cluster 1 showed higher expression of lymphoid-cell-associated genes than that of the other clusters, and cluster 3 showed higher expression of myeloid-cell-associated genes than that of the other clusters (Supplementary Fig. 5b). Cluster 2 had a mixed lympho-myeloid expression profile (Supplementary Fig. 5b). The cellular composition of each gene-expression cluster mirrored the single-cell functional output (Supplementary Fig. 5b).
a, Strategy for combining single-cell functional analysis, single-cell RNA sequencing and single-cell qRT-PCR based on flow-cytometry index data. b, Clustering of single LMPPs, GMPs and MLPs (top right key; one cell per column) by expression (bottom key; log-normalized values) of the 55 genes (right margin; one per row) with the highest and most variable expression between clusters. c,d, PCA of the 5,388 genes with highly variable expression (above the level of technical variance calculated from external control RNA), assessed by cell type (key) (c) or cluster membership (key) (d). Data are from one donor of two donors, in one experiment (other donor, Supplementary Fig. 5d–f).
We performed dimensionality reduction of gene-expression data using a diffusion- map method adapted for single cell data37,38. We indicated progenitor identity on the diffusion map (Supplementary Fig. 5c) and found that MLPs, LMPPs and GMPs cells formed a continuum, in agreement with the hierarchical clustering (Supplementary Fig. 5a). Next, we colored the diffusion map by cluster assignment (Supplementary Fig. 5c). Cluster 2 was positioned between clusters 1 and 3, in agreement with its mixed lympho-myeloid transcriptional signature (Supplementary Fig. 5b).
To overcome gene-selection bias in qRT-PCR data, we performed single-cell RNA sequencing and correlated this with function for 91 LMPPs, 110 MLPs and 119 GMPs from two different donors (157 cells and 163 cells from each donor). Clustering using the union (combined set) of variable genes (above the level of technical variance calculated from external control RNA) within each donor identified three clusters (Fig. 5b and Supplementary Fig. 5d). Most cells in cluster 1 were MLPs and most cells in cluster 3 were GMPs, while cluster 2 comprised LMPPs and GMPs (Supplementary Fig. 5e). Cluster 1 showed high expression of lymphoid-cell-associated genes (for example, MME, JCHAIN and ABCA1) (Fig. 5b and Supplementary Fig. 5d). Cluster 3 showed higher expression of myeloid genes (for example, CPA3, MPO and VIM) than that of the other clusters (Fig. 5b and Supplementary Fig. 5d). Cluster 2 showed a mixed transcriptional signature and higher expression of the hemopoietic-progenitor-cell-associated gene KIT that of the other clusters (Fig. 5b and Supplementary Fig. 5d). PCA revealed a transcriptional continuum of LMPP, MLP and GMP populations (Fig. 5c,d). Identical analysis of the second donor produced similar results (Supplementary Fig. 5f). Overall, the single-cell transcriptional profiles of the LMPPs, MLPs and GMPs suggested a continuum of lympho-myeloid differentiation for the currently defined LMPPs, MLPs and GMPs.
Refined sorting strategies purify lymphoid CD10hiCD45RAhi LMPPs and myeloid CD38hi GMPs
As our data showed that current sorting by flow cytometry does not purify functionally homogenous populations, we correlated surface-marker expression with function for the LMPPs and GMPs, as they showed the greatest functional heterogeneity. Flow-cytometry indexing data showed that single LMPPs with lymphoid output had significantly higher expression of CD10 and CD45RA than that of LMPPs with myeloid and lympho-myeloid output (Fig. 6a,b). That result was confirmed by the higher expression of CD10 in single LMPPs in the lymphoid-biased cluster 1 than in single LMPPs in the myeloid-biased cluster 3 (Supplementary Fig. 6a). Therefore, we developed a new strategy for sorting LMPPs by flow cytometry to purify CD10hiCD45RAhi LMPPs (called ‘LMPPly cells’ here) and CD10loCD45RAlo LMPPs (called ‘LMPPmix cells’ here) (Supplementary Fig. 6b), aiming to maximize the lymphoid-only potential and the mixed myeloid and lympho-myeloid potential, respectively. 26% of total LMPPs were LMPPly cells and 27% were LMPPmix cells (Fig. 6c). When cultured in SGF15/2 conditions and analyzed after 2 weeks, LMPPly cells had significantly lower cloning efficiency than that of LMPPs and LMPPmix cells but significantly higher cloning efficiency than that of MLPs (Fig. 6d and Supplementary Table 15; P < 0.0001, for all comparisons (Fisher’s exact test)). LMPPly cells were lymphoid progenitors with almost no myeloid potential and significantly lower myeloid potential than that of LMPPs or LMPPmix cells (Fig. 6d and Supplementary Table 15; P = 0.0496 and P = 0.0280, respectively (Fisher’s exact test)). LMPPly cells had very limited residual lymphoid-myeloid potential (1.6%) (Fig. 6h). LMPPmix cells retained almost all of the myeloid potential and most of the lympho-myeloid potential that the LMPPs had (Fig. 6e–h and Supplementary Table 15). This suggested that functionally, LMPPly cells were intermediate between LMPPs and MLPs. That was confirmed by a second in vitro culture (SF7b) (Fig. 6i, Supplementary Fig. 6c–f and Supplementary Table 15).
a, Expression of the surface markers CD10 and CD45RA on CB LMPPs, grouped by functional output as lymphoid (composed of uni-lymphoid output (B or NK) and bi-lymphoid output (B and NK)), myeloid (composed of uni-myeloid output (M or G) and bi-myeloid output (M and G)) or lympho-myeloid (key); expression values calculated with the ‘flowCore::logicleTransform’ function in R software. CD10, P = 0.0052, lymphoid versus lympho-myeloid, and P = 0.027, lymphoid versus myeloid; CD45RA, P = 4.8 × 10−6, lymphoid versus lympho-myeloid, and P = 0.0027, lymphoid versus myeloid (Wilcoxon rank sum test). b, Expression of CD10 and CD45RA on CB LMPPs, measured by flow cytometry (calculated as in a); colors indicate output from functional assays (key). c, Revised sorting strategy for CB cells based on expression of CD10 and CD45RA defined by bioinformatics analyses. Numbers adjacent to outlined areas indicate percent cells from the parental gate (above plots). d, Total cloning efficiency of single CB MLPs, LMPPs, LMPPly cells (56 of 244 cells), LMPPmix cells (152 of 240 cells) and GMPs in SGF15/2 culture (left), and cloning efficiency of lymphoid lineages (middle) and myeloid lineages (right) (key; presented as in Fig. 2a). e–g, Uni-lineage output e, bi-lineage output f and multi-lineage output g of single CB cells as in d (horizontal axis) in SGF15/2 culture, presented as the frequency of output cells (key) from positive wells (percent values). h,i, Lymphoid, myeloid and lympho-myeloid output (key) of CB cells as in d (horizontal axis) in SGF15/2 culture h or SF7b culture i, presented as the frequency among all plated cells. *P = 0.01–0.05, **P = 0.001–0.01 and ***P < 0.001 (Wilcoxon rank-sum test a, Fisher’s exact test (d, left) or Student’s t-test (d, middle and right)). Data are from one experiment with two donors (a,b; presented as median (middle line), lower and upper quartile (box top and bottom), minimum and maximum without outliers (bars extending above and below), and outliers (symbols above and below)), two experiments with one donor representative of six donors (c), two experiments (d, middle and right; mean + s.d. of six donors), two experiments with six donors (n = 22 donors for LMPPs, MLPs and GMPs; same donors in Fig. 2a–e) (d, left, and e–h) or two experiments with six donors (n = 9 donors for LMPPs, MLPs and GMPs, including n = 3 donors in Fig. 2f–j) (i).
On the basis of flow-cytometry indexing data, GMPs with myeloid-only output had significantly higher expression of the costimulatory molecule CD38 than that of GMPs with lympho-myeloid or lymphoid output (Fig. 7a,b). Concordantly, single GMPs in cluster 3 (with the highest myeloid potential) had significantly higher CD38 expression than that of GMPs in cluster 1 (with the highest lymphoid potential) or cluster 2 (with lymphoid and myeloid potential) (Supplementary Fig. 6g). There was a significant positive correlation between expression of CD38 and expression of a myeloid-cell-associated gene (MPO), and a negative correlation between expression of CD38 and expression of lymphoid-cell-associated genes (MME and SELL), as assessed by single cell qRT-PCR (P = 2.2 × 10−16 and ρ = 0.53 (MPO), P = 7.1 × 10−5 and ρ = –0.22 (MME), and P = 1.3 × 10−5 and ρ = –0.24 (SELL) (Spearman’s rank correlation coefficient); Supplementary Fig. 6h). To purify a GMP sub-population without lymphoid potential on the basis of CD38 expression, we divided the entire Lin–CD34+ population into CD38hi cells (44% of CD38+ cells), CD38lo cells (15% of CD38– cells) and CD38int cells (an area between the two new gates) (Fig. 7c and Supplementary Fig. 6i). We further purified CD38hi, CD38int and CD38lo cells to isolate CD38hi GMPs, CD38int (CD38intCD45RA+CD10–) cells and CD38lo LMPPs. CD38lo LMPPs were rare (1 in 1 × 108 MNCs), and no conclusions could be reached about their functional potential (data not shown). We compared the in vitro lineage potential of single CD38hi GMPs (279 cells) and CD38int cells (693 cells) with that of conventionally purified LMPPs (1,136 cells) and GMPs (1,622) using SGF15/2 culture. Whereas the CD38hi GMPs and LMPPs had a similar cloning efficiency of ~55%, the GMPs and CD38int cells had a slightly higher cloning efficiency of ~70% (Fig. 7d). All four populations produced principally uni-lineage output (63–72%) (Fig. 7e). CD38hi GMPs had much lower lymphoid potential (P < 0.0001 (Fisher’s exact test)) and lympho-myeloid potential (P = 0.0115 (Fisher’s exact test)) than that of conventionally purified GMPs (Fig. 7e–h and Supplementary Table 15), indicative of a functionally purer population. In summary, among CD10–CD45RA+ LMPPs, the phenotype CD10hiCD45RAhi marked progenitors with lymphoid-only potential, and among GMPs, the phenotype CD38hi marked progenitors with myeloid-only potential. Collectively, our single-cell analyses suggested that the progenitor hierarchy downstream of stem cells might be more complex than previous models have suggested (Supplementary Fig. 7).
a, Expression of the surface marker CD38 on CB GMPs (calculated as in Fig. 6a), grouped by functional output (key). P = 1.57 × 10−11 (myeloid versus lymphoid) and P = 1.6 × 10−8 (myeloid versus lympho-myeloid) (Wilcoxon rank-sum test). b, Expression of CD38 and CD34 by CB GMPs; colors indicate output from functional assays (key). c, Revised sorting strategy for CB cells, based on CD38 expression defined by bioinformatics analysis. Numbers adjacent to outlined areas indicate percent cells in each. d, Total cloning efficiency of single CB CD38hi GMPs (152 of 279 cells), CD38int cells (508 of 693 cells) and LMPPs (horizontal axis) in SGF15/2 culture (left), and cloning efficiency of lymphoid lineages (middle) and myeloid lineages (right) of single cells as at left (key; presented as in Fig. 2a). e–g, Uni-lineage output e, bi-lineage output f and multi-lineage output g of single CB cells as in d (horizonal axis), presented as the frequency of output cells (key) in positive wells. h, Lymphoid, myeloid and lympho-myeloid output (key) of CB cells as in d (horizonal axis), presented as the frequency among all plated cells. *P = 0.01–0.05, **P = 0.001–0.01 and ***P < 0.001 (Wilcoxon rank sum test a, Fisher’s exact test (d, left) or Students t-test (d, middle and right)). Data are from two experiments with five donors (a,b; a presented as in Fig. 6a), two experiments with one donor representative of four donors (c), two experiments (d, middle and right; mean + s.d. of four donors) or two experiments with four donors (or n = 22 donors for LMPP and GMP controls; same donors in Fig. 2a–e) (d, left, and e–h).
Discussion
Here we reported on the prospective separation and direct comparison of freshly isolated CB LMPPs, MLPs and GMPs. Our results showed that these lympho-myeloid progenitors were functionally and transcriptionally distinct and heterogeneous at the single-cell level. Although uni-lineage progenitors were most abundant, rarer multi-lineage lympho-myeloid progenitors were detected, most frequently among the LMPPs. Single-cell transcriptional analysis showed that LMPPs, MLPs and GMPs formed a transcriptional continuum. MLPs were positioned at one end of the lympho-myeloid spectrum, with GMPs at the other end, and they intersected with LMPPs, positioned in the middle. By combining functional and transcriptional analyses with flow-cytometry index data, we devised new flow-cytometry purification strategies for the isolation of myeloid CD38hi GMPs and two sub-populations within the CD10–CD45RA+ LMPP population: lymphoid CD10hiCD45RAhi (LMPPly) cells and CD10loCD45RAlo (LMPPmix) cells that retained most of the lympho-myeloid potential of LMPPs.
Several issues have prevented clear understanding of previously identified human lympho-myeloid progenitors. First, these progenitors have been isolated via cell-surface markers, on the basis of historical precedent, rather than via marker purification to functional homogeneity. Second, prospectively isolated lympho-myeloid progenitor populations have never previously been systematically compared, to our knowledge. Third, it is unclear if early progenitor populations downstream of HSCs include only uni-lineage cells16,25 or also include bi- and multi-lineage progenitors in mice5,14,17,18,26,39 or humans21,22,23,24. Fourth, functional assays demonstrate potential rather than actual cell fate in vivo under steady-state conditions. Finally, failure to register functional potential might reflect the inadequacy of an assay rather than the true potential, or indeed fate, of the cell in vivo. Thus, there is uncertainty about how distinct the differently identified progenitors are, and, if they are distinct, what their comparative functional potentials and transcriptional programs are at a clonal level.
Our exhaustive analysis of 4,598 single LMPPs, MLPs and GMPs, as well as populations of these progenitors, showed that they were functionally different in vitro and in vivo when transplanted in mice with humanized ossicles. The novel humanized ossicle model allowed ~10- to 100-fold more human-cell output than reported previously21,23,25. The GMP was mainly a progenitor of myeloid cells, with residual B cell and NK cell potential. Residual lymphoid potential was almost completely eliminated in the 44% of GMP cells with the highest CD38 expression (GMP CD38hi). The MLP was mainly a progenitor of B cells, NK cells and T cells, with residual monocyte output. The LMPP had lymphoid and myeloid potential. Our new flow-cytometry purification scheme divided the LMPPs into two populations on the basis of expression of CD10 and CD45RA: one was almost entirely lymphoid (LMPPly); the other captured most of the myeloid and lympho-myeloid potential (LMPPmix). Interestingly, the LMPPs produced mainly myeloid cells in vivo. Humanized ossicles might be particularly efficient at promoting human myelopoiesis, unlike the immunodeficient NSG mouse strain, which better supports lymphopoiesis.
We detected 24 different lineage-associated potentials in lympho-myeloid progenitors, which probably represents an underestimation, as we did not assess eosinophil, mast-cell, basophil or DC function. Although the majority of progenitors were uni-lineage, bi-lineage and multi-lineage output was seen (in up to 39% and 13%, respectively, of cells in vitro). Lympho-myeloid lineage ‘decisions’ can occur at multiple levels in HSC populations1,2,3 and MPP populations 5,14 and in presumably more-mature LMPP populations17,18,26, MLP populations and GMP populations39. Within the LMPP and GMP populations, true lympho-myeloid progenitors were rare (up to 10–14% of cells) and were concentrated among LMPPs. Notably, no experiments so far, to our knowledge, have directly assessed the hierarchical relationships among lineage-biased HSCs, MPPs and lympho-myeloid progenitors. Quantitative differences between fetal liver and BM have been observed in the multi-lineage output versus uni-lineage output of the broad CD34+CD38+ and CD34+CD38– populations16. All our data were obtained with CB, and experiments similar to those reported here will be needed to determine the proportion of uni-lineage, bi- lineage and multi-lineage progenitors in BM.
One separate question is whether diverse lineage-associated progenitors identified in vitro have stably different functions or whether there is plasticity, such that functional output might be stochastically determined or variably ‘instructed’. Further single-cell functional analysis of potentially functionally purer populations will be needed, with detailed fate mapping, in mice.
The rarity of LMPPs and MLPs (2 in 1 × 105 BM MNCs and 1 in 1 × 104 CB MNCs, respectively) and the minor proportion of multi- and bi-lineage progenitors among the LMPPs prompted us to study large numbers of single cells to obtain robust information on rare bi- and multi-lineage potentials. The rarity of LMPPs is also noteworthy for two reasons. First, single-cell RNA-sequencing analysis of unfractionated MNCs39 will require the sequencing of large numbers of cells to provide adequate representation of these rare progenitors. Second, in acute myeloid leukemia, leukemic stem cells are often arrested at an LMPP-like stage, where they can constitute up to 80% of MNCs22. Given that, we speculate that the small pool size of healthy LMPPs might be very tightly controlled to minimize oncogenic transformation. Additionally, understanding how healthy LMPPs differentiate might provide insight into novel differentiation therapies for LMPP leukemic stem cells in acute myeloid leukemia.
Methods
Collection of samples from healthy donors and patients
BM or CB samples from healthy donors were obtained with informed consent (UK protocol MREC 06/Q1606/ or Administrative Panel on Human Subjects Research Institutional Review Board-approved protocols Stanford IRB no. 18329, no. 6453, and no. 5637). Fresh CB samples were purchased from NHS Cord Blood Bank, UK or from New York Blood Center. They were processed within 16–34 h after collection. Mononuclear cells were isolated and CD34+ fraction was separated as described40. Fresh or frozen BM MNCs or CD34+ fractions were used. Human BM stromal cell were obtained from samples according to Medical University of Graz Ethikkommission (Institutional Review Board-approved protocol, MUG Graz IRB no. 19-252). BM mesenchymal stromal cells (MSCs) were isolated and cell populations were expanded as described34.
Sorting of HSPC populations by flow cytometry
Antibodies used for sorting by flow-cytometry and immunophenotyping are listed in Supplementary Table 16. CB or BM MNCs enriched for CD34+ cells underwent depletion of lineage-marker-positive cells by staining with purified antibodies to human CD2, CD3, CD4, CD7, CD8a, CD11b, CD14, CD19, CD20, CD56 and CD235a, followed by Qdot 605 conjugated goat F(ab’)2 anti-mouse IgG (H + L). Cells were also stained for human markers with anti-CD38-FITC, anti-CD45RA-PE or anti-CD45RA-BV650, anti-CD123-PE Cy7, anti-CD90-biotin, anti-CD34-PerCP and anti-CD10-APC. Finally, cells were incubated with streptavidin-conjugated APC-eF780 and Hoechst 33258 (Invitrogen, Loughborough UK; final concentration: 1 μg/ml). For the humanized-ossicle-xenotransplantation assay, CD34+ CB cells were stained with purified antibodies to the human lineage markers CD2, CD3, CD4, CD7, CD8a, CD11b, CD14, CD19, CD20, CD56, CD235a and anti-CD16. All antibodies to lineage markers were PE Cy5 conjugated. Cells were then stained with anti-CD38-PE Cy7, anti-CD90-FITC, anti-CD123-PE, anti-CD34-APC, anti-CD10-APC Cy7, anti-CD45RA-BV605 and propidium-iodide (Thermo Fisher, Waltham MA; final concentration: 1 μg/ml). Unstained, single-stained and fluorescence-minus-one (FMO) controls were used to determine background staining and compensation in each channel. Single-stained controls used the anti-mouse Ig, κ/Negative Control compensation particle set (BD, Oxford UK). CB cells were sorted with an average purity of 99% for in vitro and RNA assays and 96% for humanized ossicle xenotransplantation. Prior to single-cell sorting, single fluorescent beads were deposited directly into a 96-well plate to establish accuracy of single-cell deposition (>99%). Sorting was performed on a BD FACSAria III or BD Fusion, and flow-cytometry analysis was done on LSR Fortessa X20. Data analysis was performed using Diva v8.1 or FlowJo v10.0.06 and v10.0.07r2.
Index sorting for functional and transcriptional analyses
For index sorting, we saved information on the following parameters: FSC, SSC, Hoechst and expression of lineage markers, CD34, CD38, CD45RA, CD10, CD90 and CD123 for each single cell. For 919 index-sorted single cells, we assessed expression of 96 genes by qRT-PCR (Supplementary Fig. 5); 74 passed quality control. Separately, we performed single-cell index sorting and single-cell in vitro functional assays of 3,458 single cells (from Fig. 2, Supplementary Fig. 2, Fig. 6, Fig. 7 and Supplementary Fig. 6). In separate experiments, we index sorted 320 single cells for single-cell RNA-seq (Fig. 5). Using common ‘position of the cells’ in flow-cytometry plots, we could then map functional potential (i.e., lymphoid, myeloid or lympho-myeloid) to gene expression, cell-surface-marker expression and forward and side scatter. To purify LMPPly and LMPPmix cells, the thresholds were defined based on maximum expression of CD10 and CD45RA by LMPPs with myeloid output. To purify CD38hi GMPs, thresholds were set using the maximum normalized CD38 expression of GMPs with lymphoid and lympho-myeloid output.
In vitro lympho-myeloid differentiation assay (bulk, single-cell, limiting-dilution assay)
For population analysis, MS-5 cells were seeded on a 24-well plate coated with 0.1% gelatin at a density of 2 × 104 cells per well in α-MEM medium (Gibco/Thermo Fisher Scientific Loughborough UK) supplemented with 10% FBS (Hyclone, GE Healthcare, SH30070.03 Amersham Hatfield, UK), 1% penicillin-streptomycin, 1% l-glutamine, 0.1 μM DuP-697 (Cayman Chemical, Ann Arbor, Mi, USA), 20 ng/ml SCF, 10 ng/ml G-CSF, 10 ng/ml FLT3L, 10 ng/ml IL-15 and 10 ng/ml IL-2 (Peprotech London UK, SGF15/2 culture). 24 h after plating of MS-5 cells, 150 highly purified LMPPs, MLPs or GMPs were deposited into each well. Half of the medium was changed every week. Harvested cells were analyzed by flow cytometry at weeks 1, 2 and 3.
The limiting-dilution assay (LDA) was performed by sorting of LMPPs, MLPs or GMPs at different cell doses (1, 2, 5, 10 and 20 cells) from four different CB samples into 96-well plates pre-plated with 2,500 MS-5 cells per well with 100 µl of medium without cytokines. Immediately after sorting, 100 µl of 2 × SGF15/2 culture medium was added to each well. Half of the medium was changed every week. A total of 833 LMPPs, 789 MLPs and 1252 GMPs from four different CB samples were analyzed for the LDA at weeks 2–2.5 (Supplementary Table 2a). Frequency calculations were performed using L-Calc software (Stem Cell Technologies) and were independently verified by ELDA software (http://bioinf.wehi.edu.au/software/elda/). The LDA plots were generated using R, with lines representing the estimates calculated by ELDA software.
For single-cell analysis, single LMPPs, MLPs and GMPs were deposited into 96-well plates pre-plated with 2,500 MS-5 cells per well with 100 µl of medium without cytokines. Medium with 2 × cytokines was added to each well after sorting. Half of the medium was changed every week. After culture for 2–2.5 weeks, flow cytometry was performed, and wells with more than 15 human CD15+, CD14+, CD56+ or CD19+ cells were considered positive (details in Supplementary Table 2b). To compare with published conditions21, single LMPPs, MLPs and GMPs were cultured for 4 weeks on MS-5 stroma in H5100 medium (StemCell Technologies Cambridge UK) supplemented with 100 ng/ml SCF, 20 ng/ml IL-7, 50 ng/ml thrombopoietin, 10 ng/ml IL-2, 20 ng/ml GM-CSF, 20 ng/ml G-CSF and 10 ng/ml M-CSF (all from Peprotech, S7T2GM/G/M culture) and were analyzed by flow cytometry.
To read lineage readouts for all in vitro lympho-myeloid differentiation assays, harvested cells were stained with anti-human CD15-FITC, anti-human CD14-PE, anti-human CD19-PE Cy7, anti-human CD56-APC or -PE Cy5, anti-human CD45-APC Cy7 and, in some cases, with anti-human CD34-BV605 (Supplementary Table 16).
In vitro T cell–differentiation assay
OP9-hDL1 cells41 were seeded on a 24-well plate coated with 0.1% gelatin at a density of 2 × 104 cells per well in freshly prepared α-MEM medium (Gibco/Thermo Fisher Scientific, 12000-063) with 20% heat-inactivated FBS (Hyclone, GE Healthcare, SH30070.03 Amersham Hatfield, UK), 1% penicillin-streptomycin, 1% l-glutamine. 24 h after plating of OP9-hDL1 cells, 150 highly purified LMPPs, MLPs or GMPs were deposited into each well and were cultured in the presence of 10 ng/ml SCF, 5 ng/ml FLT3L and 5 ng/ml IL-7 (Peprotech, London, UK, SF7a culture). Cells were dissociated from wells and were transferred to new plates with fresh OP9-hDL1 cells weekly. Harvested cells were analyzed by flow cytometry at weeks 4, 5 and 7. Cells were stained with anti-human CD7-FITC, anti-human CD1a-PE, anti-human CD8-PE Cy7, anti-human CD4-APC and anti-human CD45-APC Cy7 (Supplementary Table 16).
In vitro combined T cell–lympho-myeloid differentiation assay
MS5-hDL1IND100 cells42 (in which hDL1 expression could be induced by the addition of doxycycline) were seeded on 96-well plates coated with 0.1% gelatin at a density of 2,500 cells per well in 100 µl freshly prepared α-MEM medium (Gibco/Thermo Fisher Scientific, Loughborough UK) supplemented with 20% FBS (Hyclone, GE Healthcare, SH30070.03HI, Amersham Hatfield, UK), 1% penicillin-streptomycin and 1% l-glutamine. 24 h after plating of MS5-hDL1IND cells, single LMPPs, MLPs or GMPs were deposited into each well and were cultured in the presence of 20 nM insulin (Sigma-Aldrich, St Louis, MO), 50 ng/ml SCF, 20 ng/ml FLT3L and 10 ng/ml IL-7 (Peprotech London UK, SF7b culture). Fresh medium was added every week.
Cells were harvested at 21 d and were split into two groups: half were used for flow cytometry, and the remaining half were re-seeded onto MS5-hDL1IND100 cells and cultured in SF7b/Dox culture with doxycycline (1 μg/ml). Half of the medium was changed twice every week. Fresh doxycycline was added to the cultures three times a week. At 42 d, cells were harvested and flow cytometry was performed. At 21 d, wells with more than 8 human CD15+, CD14+, CD56+ or CD19+ cells were considered positive. At 42 d, flow cytometry with anti-human CD1a, anti-human CD7, anti-human CD4 and anti-human CD8 was performed, and wells with more than 8 CD7+ cells were considered positive for T cell output.
CFU assay
Colony formation was tested as before22. Colony identity was confirmed morphologically after cytospin (medium acceleration, 800 r.p.m., 5 min, May–Grunwald–Giemsa stain; Sigma, Poole UK) and by flow cytometry with anti-human CD15-FITC, anti-human CD14-PE and anti-human CD235a-PE Cy5 (Supplementary Table 16).
Humanized-ossicle-xenotransplantation assay
Protocol was performed as previously described34. In brief, in vitro–expanded human BM-MSC populations were harvested, resuspended in 60 μl of pooled human platelet lysate (pHPL) and admixed with 240 μl of matrigel-equivalent matrix. The whole matrix-cell mixture was injected subcutaneously to generate humanized ossicle niches. 8–10 weeks after application of BM-MSCs, transplants were evaluated for bone and marrow formation. Mice with established humanized ossicle niches were conditioned with 200 rads of irradiation 12–24 h before transplantation. Variable numbers of LMPPs, MLPs and GMPs from at least three different CB donors (Supplementary Fig. 3c) were transplanted in total volume of 20 μl by direct intraossicle injection. Experiments were performed in accordance with a protocol approved by Stanford’s Administrative Panel on Laboratory Animal Care (no. 22264) and in adherence to the US National Institutes of Health’s Guide for the Care and Use of Laboratory Animals. Normal multi-lineage engraftment was assessed 1–2 weeks after transplantation and was defined by the presence of myeloid cells (CD33+) and B cells (CD19+) among engrafted human CD45+HLA-ABC+ cells. Engrafted mice were stained with anti-human CD14-PE or anti-human CD14-APC Cy7, anti-human CD15-FITC, anti-human HLA-ABC-FITC or anti-human HLA-ABC-PB, anti-human CD19-APC, anti-human CD33-PE and anti-human CD45-V450 (Supplementary Table 16).
RNA sequencing of bulk HSPC populations
100 highly purified HSPCs from CB samples obtained from healthy donors were sorted directly into lysis buffer containing RNAse inhibitor (Clontech St Germain-en-Laye France) and were stored at –80°C before further processing. cDNA synthesis was done with Smarter Ultra low input RNA kit v1 (Clontech) as previously described43. Illumina libraries were generated using a Nextera XT DNA sample preparation kit and Index Kit (Illumina Chesterford UK). Library size and quality were checked using Agilent High-Sensitivity DNA chip with Agilent Bioanalyser (Agilent Technologies Stockport UK). The concentration of indexed libraries was determined using a Qubit High-Sensitivity DNA kit (Invitrogen Loughborough, UK). Libraries were pooled to a final concentration of 5–14 nM and were sequenced on an Illumina HiSeq 2000 single-end 50-bp reads.
Bulk and single-cell gene-expression analysis by Dynamic Arrays
Gene-expression analysis was performed as described40. TaqMan assays (Applied Biosystems) are listed in Supplementary Tables 2 and 8.
Single-cell RNA sequencing
Single-cell libraries for RNA sequencing were prepared using the Smart-seq2 protocol44, whereby 23 cycles were used for the cDNA library preamplification. Illumina Nextera XT DNA sample preparation kit and Index Kit (Illumina Chesterford UK) was used for cDNA tagmentation and indexing. ERCC RNA Spike-In Mix (Ambion) was added to the lysis mix at a final dilution of 1:80,000,000. Library size, quality and concentration were checked as done for the bulk RNA sequencing. Libraries were pooled to a final concentration of 7–28 nM, and 78–95 single-cell libraries were combined per pool. Sequencing was done on HiSeq4000 using 75-bp paired-end reads. Each pool contained a library generated from an empty well.
Bioinformatics analysis (bulk RNA-seq, single-cell Biomark and single-cell RNA-seq)
For 50-bp single-end bulk RNA sequencing, alignment to the hg38 reference genome was carried out using TopHat v2.0.10 (ref. 45). Alignments were processed using Picard tools (http://picard.sourceforge.net/). We used R version 3.1.1 http://www.R-project.org. Sequencing reads were filtered for mapq 4 i.e. uniquely mapping reads. This gave a range of 15.1 × 106 to 56.2 × 106 aligned reads. The total number of genes expressed per sample was calculated as an rpkm >1. The number of expressed genes ranged from 7,707 to 11,350, with an average of 9,800. The count matrix was transformed to log2(cpm) scale, and principal-component analysis was carried out. An ANOVA-like test was performed, using edgeR package for R, to identify genes expressed differentially by the populations. One CB biological replicate MPP population was excluded because its global gene expression showed more uniquely expressed genes than in the three remaining MPP biological replicates and showed low correlation to the other three replicates. The genes were ranked by their significance (P value adjusted for multiple testing), and different numbers of genes were used for PCA and hierarchical clustering of samples. Eigenvalues from PCA were calculated by using the square of the s.d. of the principle components. Differential gene expression for one versus one and one versus all comparisons were calculated using edgeR. For gene-signature generation, a cut-off of a change in expression of over onefold (log values) was used, and genes were ranked by P value. Heat maps and associated hierarchical clustering were generated using GENE-E software (Broad Institute) or using the R packages pvclust and heatmap.2 (gplots). MetaCore Pathway Map enrichment analysis (Thomson Reuters, London UK) was carried out on genes expressed differentially by each lympho-myeloid population versus all other populations (one versus all). A P-value cut-off of 0.05 was used to identify positively enriched pathway maps.
Analysis of single-cell Biomark data was performed in R version 3.3.1 using data exported from the Fluidigm Data Collection software. For quality control, amplification curves with a Quality Score of <0.65, and any Ct values of >27 were treated as undetected expression. Any cells in which expression of both housekeeping genes (B2M and GAPDH) was not detected were removed from further analysis (n = 7 cells). An additional cell was removed as it had a large outlying number of genes detected. The housekeeping gene ACTB was also measured in the assay, but unlike B2M and GAPDH, it did not show robust expression across the majority of cells and therefore was not used in further analysis. Normalized ΔCt values were calculated by subtraction of the mean of Ct values for B2M and GAPDH in each cell, as previously described19. Housekeeping genes were excluded from further analysis. Genes detected in <20 cells, with a variance of <1 across all cells or expressed in none of the MLP, GMP or LMPP ten-cell control samples assayed by qRT-PCR alongside single-cell samples were removed from downstream analysis. Post-quality-control data measured 74 genes in 919 single cells.
Hierarchical clustering was performed on genes and cells by using the hclust function (stats package) with distance measure 1 – Spearman’s correlation and agglomeration method Ward.D2. The heat map visualizing the clustering was plotted using the heatmap.2 function (gplots package). Cells were divided into three clusters using the cutree function (stats package) on the hierarchical clustering. A gene was classed as expressed differentially by two clusters if it satisfied two criteria: 1) the magnitude of the change (log2 fold value) of the mean ΔCt in each cluster was >1, and 2) the adjusted P value (Benjamini & Hochberg correction for multiple testing) of two-sided Wilcox test of ΔCt expression values between the two clusters was <0.01. Diffusion maps46 were used for dimensionality reduction of the single-cell gene-expression data. This method was implemented using the DiffusionMap function from the destiny R package with Euclidean distance37,47.
Single-cell RNA-sequencing reads were aligned using G-SNAP48, and mapped reads were assigned to Ensembl genes (release 81; ref. 49) by using HTSeq50. Cells with fewer than 500,000 reads mapping to nuclear genes, greater than 20% of mapped reads mapping to mitochondrial genes or greater than 20% of mapped reads mapping to External RNA Controls Consortium (ERCC) spike-ins or with expression of fewer than 750 different genes with at least ten counts were removed from further analysis. ERCC spike-in controls identified genes exceeding technical variance51. From donors 1 and 2, 163 of 166 cells and 157 of 249 cells, respectively, passed quality control. Single-cell profiles were normalized using the scran R package52, and variable genes were identified as having variation exceeding technical levels51. Data showed batch effects between different donors. The Seurat R package (https://github.com/satijalab/seurat) was then used to regress out plate effects from the sequencing data and set more stringent thresholds for variable genes, leading to 1,605 variable genes in donor 1 and 1,273 variable genes in donor 2. Principal-component analysis was performed using Seurat, and clusters were found using the Seurat::FindClusters function on the first ten principal components. Heat maps display the top genes marking these clusters as identified by the Seurat::FindAllMarkers function and were visualized using the gplots::heatmap.2 function.
Statistical analysis
Frequency of populations in flow-cytometry plots gates is the mean of the population across all samples analyzed as indicated. Bar graphs of gene-expression analysis represent mean + s.e.m. or mean + s.d., as indicated in figure legends. Two-tailed Student’s unpaired t-test and Fisher’s exact test (Excel, GraphPad sortware) were used to determine statistical significance in gene-expression-analysis data and single-cell functional assays, respectively. The statistical significance of the P value was defined as follows for all P value comparisons made: P > 0.05. not significant; P = 0.01–0.05, significant (*), P = 0.001–0.01, very significant (**); and P < 0.001, extremely significant (***). Wilcoxon rank sum test was done using R. Kruskal-Wallis test, stratified by group, was used to define significant differences between LMPPs, MLPs and GMPs in the single-cell functional assay in SGF15/2 culture and gave the following P values: LMPPs, 5 × 10−6; MLPs, 0.1725; and GMPs, 0.7395. Wilcoxon rank sum test confirmed that there were no outliers among single LMPPs from different CB donors. Prism software was used to plot the gene-expression analysis and single-cell in vitro data. LDA plots were generated using R, and the lines represent the estimates calculated using ELDA software.
Life Sciences Reporting Summary
Further information on experimental design and reagents is available in the Life Sciences Reporting Summary.
Data availability
Bulk RNA sequencing data have been deposited in Arrayexpress (https://www.ebi.ac.uk/arrayexpress/) with accession number E-MTAB-5456. The single-cell RNA-sequencing data accession number is GSE100618. All other source data that support the findings of this study are available from the corresponding author upon request.
References
Dykstra, B. et al. Long-term propagation of distinct hematopoietic differentiation programs in vivo. Cell Stem Cell 1, 218–229 (2007).
Challen, G. A., Boles, N. C., Chambers, S. M. & Goodell, M. A. Distinct hematopoietic stem cell subtypes are differentially regulated by TGF-beta1. Cell Stem Cell 6, 265–278 (2010).
Benz, C. et al. Hematopoietic stem cell subtypes expand differentially during development and display distinct lymphopoietic programs. Cell Stem Cell 10, 273–283 (2012).
Chen, J. Y. et al. Hoxb5 marks long-term haematopoietic stem cells and reveals a homogenous perivascular niche. Nature 530, 223–227 (2016).
Oguro, H., Ding, L. & Morrison, S. J. SLAM family markers resolve functionally distinct subpopulations of hematopoietic stem cells and multipotent progenitors. Cell Stem Cell 13, 102–116 (2013).
Sanjuan-Pla, A. et al. Platelet-biased stem cells reside at the apex of the haematopoietic stem-cell hierarchy. Nature 502, 232–236 (2013).
Yamamoto, R. et al. Clonal analysis unveils self-renewing lineage-restricted progenitors generated directly from hematopoietic stem cells. Cell 154, 1112–1126 (2013).
Sun, J. et al. Clonal dynamics of native haematopoiesis. Nature 514, 322–327 (2014).
Busch, K. et al. Fundamental properties of unperturbed haematopoiesis from stem cells in vivo. Nature 518, 542–546 (2015).
Sawai, C. M. et al. Hematopoietic stem cells are the major source of multilineage hematopoiesis in adult animals. Immunity 45, 597–609 (2016).
Yu, V. W. et al. Epigenetic memory underlies cell-autonomous heterogeneous behavior of hematopoietic stem cells. Cell 167, 1310–1322 (2016).
Wilson, A. et al. Hematopoietic stem cells reversibly switch from dormancy to self-renewal during homeostasis and repair. Cell 135, 1118–1129 (2008).
Cabezas-Wallscheid, N. et al. Identification of regulatory networks in HSCs and their immediate progeny via integrated proteome, transcriptome, and DNA methylome analysis. Cell Stem Cell 15, 507–522 (2014).
Pietras, E. M. et al. Functionally distinct subsets of lineage-biased multipotent progenitors control blood production in normal and regenerative conditions. Cell Stem Cell 17, 35–46 (2015).
Majeti, R., Park, C. Y. & Weissman, I. L. Identification of a hierarchy of multipotent hematopoietic progenitors in human cord blood. Cell Stem Cell 1, 635–645 (2007).
Notta, F. et al. Distinct routes of lineage development reshape the human blood hierarchy across ontogeny. Science 351, aab2116 (2016).
Adolfsson, J. et al. Identification of Flt3+ lympho-myeloid stem cells lacking erythro-megakaryocytic potential a revised road map for adult blood lineage commitment. Cell 121, 295–306 (2005).
Månsson, R. et al. Molecular evidence for hierarchical transcriptional lineage priming in fetal and adult stem cells and multipotent progenitors. Immunity 26, 407–419 (2007).
Guo, G. et al. Mapping cellular hierarchy by single-cell analysis of the cell surface repertoire. Cell Stem Cell 13, 492–505 (2013).
Naik, S. H. et al. Diverse and heritable lineage imprinting of early haematopoietic progenitors. Nature 496, 229–232 (2013).
Doulatov, S. et al. Revised map of the human progenitor hierarchy shows the origin of macrophages and dendritic cells in early lymphoid development. Nat. Immunol. 11, 585–593 (2010).
Goardon, N. et al. Coexistence of LMPP-like and GMP-like leukemia stem cells in acute myeloid leukemia. Cancer Cell 19, 138–152 (2011).
Kohn, L. A. et al. Lymphoid priming in human bone marrow begins before expression of CD10 with upregulation of L-selectin. Nat. Immunol. 13, 963–971 (2012).
Görgens, A. et al. Multipotent hematopoietic progenitors divide asymmetrically to create progenitors of the lymphomyeloid and erythromyeloid lineages. Stem Cell Rep. 5, 154–155 (2015).
Velten, L. et al. Human haematopoietic stem cell lineage commitment is a continuous process. Nat. Cell Biol. 19, 271–281 (2017).
Perié, L., Duffy, K. R., Kok, L., de Boer, R. J. & Schumacher, T. N. The branching point in erythro-myeloid differentiation. Cell 163, 1655–1662 (2015).
Berardi, A. C. et al. Individual CD34+CD38lowCD19–CD10– progenitor cells from human cord blood generate B lymphocytes and granulocytes. Blood 89, 3554–3564 (1997).
Ichii, M. et al. The density of CD10 corresponds to commitment and progression in the human B lymphoid lineage. PLoS One 5, e12954 (2010).
Farlik, M. et al. DNA methylation dynamics of human hematopoietic stem cell differentiation. Cell Stem Cell 19, 808–822 (2016).
Lee, J. et al. Restricted dendritic cell and monocyte progenitors in human cord blood and bone marrow. J. Exp. Med. 212, 385–399 (2015).
Pronk, C. J. et al. Elucidation of the phenotypic, functional, and molecular topography of a myeloerythroid progenitor cell hierarchy. Cell Stem Cell 1, 428–442 (2007).
Galy, A., Travis, M., Cen, D., Chen, B. & Human, T. Human T, B, natural killer, and dendritic cells arise from a common bone marrow progenitor cell subset. Immunity 3, 459–473 (1995).
Manz, M. G., Miyamoto, T., Akashi, K. & Weissman, I. L. Prospective isolation of human clonogenic common myeloid progenitors. Proc. Natl. Acad. Sci. USA 99, 11872–11877 (2002).
Reinisch, A. et al. A humanized bone marrow ossicle xenotransplantation model enables improved engraftment of healthy and leukemic human hematopoietic cells. Nat. Med. 22, 812–821 (2016).
Novershtern, N. et al. Densely interconnected transcriptional circuits control cell states in human hematopoiesis. Cell 144, 296–309 (2011).
Wilson, N. K. et al. Combined single-cell functional and gene expression analysis resolves heterogeneity within stem cell populations. Cell Stem Cell 16, 712–724 (2015).
Haghverdi, L., Buettner, F. & Theis, F. J. Diffusion maps for high-dimensional single-cell analysis of differentiation data. Bioinformatics 31, 2989–2998 (2015).
Moignard, V. et al. Decoding the regulatory network of early blood development from single-cell gene expression measurements. Nat. Biotechnol. 33, 269–276 (2015).
Paul, F. et al. Transcriptional heterogeneity and lineage commitment in myeloid progenitors. Cell 163, 1663–1677 (2015).
Quek, L. et al. Genetically distinct leukemic stem cells in human CD34– acute myeloid leukemia are arrested at a hemopoietic precursor-like stage. J. Exp. Med. 213, 1513–1535 (2016).
Six, E. M. et al. Cytokines and culture medium have a major impact on human in vitro T-cell differentiation. Blood Cells Mol. Dis. 47, 72–78 (2011).
Calvo, J., BenYoucef, A., Baijer, J., Rouyez, M. C. & Pflumio, F. Assessment of human multi-potent hematopoietic stem/progenitor cell potential using a single in vitro screening system. PLoS One 7, e50495 (2012).
Woll, P. S. et al. Myelodysplastic syndromes are propagated by rare and distinct human cancer stem cells in vivo. Cancer Cell 25, 794–808 (2014).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Coifman, R. R. et al. Geometric diffusions as a tool for harmonic analysis and structure definition of data: diffusion maps. Proc. Natl. Acad. Sci. USA 102, 7426–7431 (2005).
Angerer, P. et al. destiny: diffusion maps for large-scale single-cell data in R. Bioinformatics 32, 1241–1243 (2016).
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010).
Yates, A. et al. Ensembl 2016. Nucleic Acids Res. 44 D1, D710–D716 (2016).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013).
Lun, A. T., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75(2016).
Acknowledgements
We thank J.C. Zuniga-Pflucker for OP9-DL4 cells used for initial experiments; and the High-Throughput Genomics Group at the Wellcome Trust Centre for Human Genetics (funded by Wellcome Trust (090532/Z/09/Z)) for generation of sequencing data. Supported by the MRC (MHU Award G1000729 and MRC Disease Team Award 4050189188 to P.V.; PhD studentship to Z.A. and F.H.; Single Cell Award MR/M00919X/1), CRUK (Program Grant C7893/A12796 to P.V.; Program Grant C1163/A21762 to B.G.; Development Fund CRUKDF0176-DK to D.K. and P.V.; and CRUK Development Fund C5255/A20758 to B.S. and P.V.), Bloodwise (Specialist Program 13001 and Project Grant 12019), the Oxford Partnership Comprehensive Biomedical Research Centre (NIHR BRC Funding scheme), the US National Institutes of Health (R01CA188055 and U01HL099999 to R.M.), the New York Stem Cell Foundation (Robertson Investigator, R.M.), the Leukemia and Lymphoma Society (Scholar Award to R.M.) and the Austrian Science Fund (FWF) (Erwin-Schroedinger Research fellowship to A.R.).
Author information
Authors and Affiliations
Contributions
D.K. and B.S. contributed equally to this work and are listed alphabetically; F.H. and A.R. contributed equally to this work; D.K., B.S., Z.A. and P.V. designed the experiments; D.K., B.S., Z.A., A.R., M.S., L.Q. and N.G. performed experiments and analyzed data; F.H., G.O., Z.A., E.R. and S.T. performed bioinformatics and statistical analysis; J.D. and B.U. prepared samples; J.C., E.S., F.P., C.P., R.M. and B.G. provided reagents and materials; D.K., B.S. and P.V. wrote the paper; and all authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Immunophenotypic separation of eight distinct human hemopoietic stem–progenitor cell (HSPC) populations and in vitro functional lympho-myeloid potential of LMPPs, MLPs and GMPs
(a) Representative plot of flow sorting of human CB HSPCs. Population frequencies shown are the mean from 44 CB donors calculated as a percentage of Lin-CD34+ compartment. HSC, hemopoietic stem cell; MPP, multipotent progenitor; LMPP, lymphoid-primed multi-potential progenitor; MLP, multi-lymphoid progenitor; CMP, common myeloid progenitor; GMP, granulocyte-monocyte progenitor; MEP, megakaryocyte-erythroid progenitor; B/NK, B-NK cell progenitor. (b) Representative flow cytometric analysis of the B cell, NK, monocytic and granulocytic output of CB LMPP, MLP and GMP cultured for 2 weeks on SGF15/2. Frequencies shown are mean from 3 CB donors calculated as a percentage of human CD45+ cells. (c) Representative flow cytometric analysis of T cell output of CB LMPP, MLP and GMP cultured for 5 weeks on SF7a. Frequencies shown are average from 5 CB donors calculated as a percentage of human CD45+ cells.
Supplementary Figure 2 LMPPs and GMPs are lympho-myeloid progenitors, while MLPs are mainly lymphoid progenitors in quantitative in vitro assays
(a) Experimental strategy for sorting, culture and analysis for quantitative in vitro assays in SGF15/2. Similar strategy was used for the other culture conditions. (b) LDA and single cell in vitro culture conditions used in the study. (c) Representative flow cytometric profiles of the outputs from the quantitative in vitro assays. Plots are gated on human CD45+ cells. (d) LDA plots showing the frequency (f: 1 in X cells can give rise to) of LMPP, MLP and GMP cells with B-cell, NK cell, monocytic and granulocytic potential. Plots are generated in R and the lines represent the estimates calculated using ELDA software. (e) Total cloning efficiency (left) of single LMPP, MLP and GMP in S7T2GM/G/M culture (LMPP: 69/96 cells, MLP: 4/52, GMP: 98/110). Significance defined using Fisher’s exact test. Cloning efficiency of lymphoid (Ly, middle) and myeloid lineages (My, right). Bars indicate total cloning efficiency; filled portion indicates the proportion of lymphoid (lymphoid plus mixed clones) or myeloid potential (myeloid plus mixed clones). Mean ± SD is shown. Significance is defined using student’s t-test. (f) Single-, (g) bi- and (h) multi-lineage single cell outputs in S7T2GM/G/M culture, presented as percentage of the positive wells. (i) Lymphoid (Ly), myeloid (My) and lympho-myeloid (Ly-My) outputs presented as percentage of all plated single cell LMPP, MLP and GMP wells in S7T2GM/G/M culture. (j) Summary of lineage outputs from single LMPP, MLP and GMP. For the LDA data were obtained from 4 CB donors, for the single cell assay in S7T2GM/G/M culture data were obtained from 2 CB donors.
Supplementary Figure 3 Human LMPPs, MLPs and GMPs have distinct differentiation potential in vivo
(a) Experimental strategy for generation of human ossicles in NSG mice and human progenitor transplantation. (b) Gating strategy for the sorting of human progenitors for in vivo transplantation. (c) Number of cells of LMPP, MLP and GMP populations injected per ossicle. (d) Correlation between the number of transplanted cells and the myeloid/lymphoid ratio of the output cells.
Supplementary Figure 4 Transcriptional profiling of human HSPCs in bulk shows the distinct transcriptional patterns of human progenitor populations
(a) Heatmap showing hierarchical clustering and the expression of all genes by HSPC populations. Expression values are normalized per gene (per row). (b) Plot comparing the eigenvalues for each principle component (PC) for the PC analysis (PCA) of HSPC using the top 300 ANOVA genes (black) and 300 randomly selected genes from a randomized expression matrix (red). (c) 3D PCA plot showing HSPC populations using the top 300 ANOVA genes. Percentage variance for each PC is shown. (d) PCA plots showing HSPC populations using between 1,000 and 10,000 genes with the highest variance across all HSPC populations. Percentage variance for each PC is shown. (e) Table of differentially expressed genes in one versus one comparisons of HSPCs. Genes are upregulated in a population column versus row. (f) Heatmap showing the expression of genes recently identified as being expressed by lineage primed hemopoietic cells25. Genes are color-coded according to their classification. Expression values are normalized per gene. (g-h) Heatmap showing the expression of (g) hemopoietic-related transcription factors that were differentially expressed between the MLP and GMP and (h) cytokine and chemokine related genes by the LMPP, GMP and MLP. Genes affiliated with the lymphoid or myeloid lineages are color-coded (lymphoid: orange, myeloid: green) and genes associated with immune function are labeled in black. Expression values are normalized per gene.
Supplementary Figure 5 Transcriptional heterogeneity of single lympho-myeloid progenitor cells (a) Heatmap showing hierarchical clustering on genes (rows) and single cells (columns), based on qRT-PCR gene expression analysis
(b) Top, cell type composition of the 3 clusters identified in the dendrogram in (a). Middle, differentially expressed genes between cluster pairs. Bottom, expected number of cells per cluster, based on the functional assays. (c) Diffusion map dimensionality reductions colored in by cell type (left) and cluster membership (right), based on qRT-PCR gene expression analysis. (d) Heatmap showing clustering of single LMPP, GMP and MLP using the 55 most highly and variably expressed genes between clusters. Single cell RNA-sequencing data from two donors were used to generate the clusters. The heat map shows clustering from one of two. Data from the other donor is in Fig. 5b. Log-normalised gene expression (rows) for each single cell (columns) is shown. (e) Cell type composition of the 3 clusters identified in the heatmap in (d) and Fig. 5b. (f) PCA plot colored by cell type (top) or cluster membership (bottom). For single cell qRT-PCR gene expression analysis: data from 4 CB donors. For single cell RNA sequencing analysis: data from 2 donors (donor 1 in Fig. 5, donor 2 in current figure).
Supplementary Figure 6 Further functional purification of the lympho-myeloid progenitor populations (a) CD10 fluorescence intensity in LMPPs in the 3 clusters identified from clustering of single cell RNA Seq data presented in . Wilcoxon rank sum test is used to define significance.
(b) Histograms showing expression of CD10 and CD45RA for LMPP with different functional output. The percentage cut-offs used to define the new sorting strategy for LMPPly and LMPPmix are shown. Thresholds were defined based on maximum CD10 and CD45RA expression of LMPPs with myeloid output. (c) Total cloning efficiency (left) of single MLP, LMPP, LMPPly, LMPPmix and GMP in SF7b culture (LMPPly: 26/108 cells, LMPPmix: 48/100). Significance defined using Fisher’s exact test. Cloning efficiency of lymphoid (Ly, middle) and myeloid lineages (My, right). Bars indicate total cloning efficiency; filled portion indicates the proportion of lymphoid (lymphoid plus mixed clones) or myeloid potential (myeloid plus mixed clones). Mean ± SD is shown. Significance is defined using students t-test. (d) Single-, (e) bi- and (f) multi-lineage outputs from single cells in SF7b culture, presented as percentage of the positive wells. (g) CD38 fluorescence intensity in GMPs in the 3 clusters identified from clustering of single cell RNA Seq data presented in Fig. 5b. Wilcoxon rank sum test is used to define significance. (h) Correlation between expression of selected genes and cell surface marker expression in GMPs. ρ indicates Spearman’s rank correlation coefficient values and p is corresponding p-value from cor.test function in R. (i) Histograms showing expression of CD38 for GMP with different functional output. The percentage cut-off used to define the new sorting strategy for GMP CD38hi and CD38mid are shown. For functional assay based on the revised LMPP sorting strategy (SF7b culture) data are from 6 CB donors; for LMPP, MLP and GMP controls data from 9 CB donors (including 3 CB donors used for Fig. 2f-j).
Supplementary Figure 7 Models of human lympho-myeloid differentiation
(a) Lineage output of lympho-myeloid progenitors (b-c) Models of human hemopoietic differentiation and lineage diversification. Dashed arrows - hierarchical relationships not experimentally validated. (d) Novel model of human lympho-myeloid differentiation. Multiple, rare, functionally distinct bi- and multi-lineage lympho-myeloid progenitors (LMPs) could differentiate either directly from hemopoietic stem cells (HSC) or multi-potent progenitors (MPP). We have used the term LMP, rather than LMPP (lymphoid primed multi-potential progenitor) to describe these lympho-myeloid progenitors, as not all LMPs will be lymphoid biased. Multi-lineage LMPs are rare. Lymphoid only and myeloid only progenitors are shown below. Bi-lineage progenitors are more frequent and uni-potent progenitors are most common. The hierarchical relationships between LMP and lymphoid and myeloid only bi- and uni-lineage progenitors remain to be determined. This model also leaves open the question of when commitment to either lymphoid or myeloid fate occurs. Erythro-megakaryocytic differentiation leading to a MEP (megakaryocyte-erythroid progenitor), erythroid (BFU-E) and megakaryocytic progenitor (MkP) could occur directly from either HSC or MPP.
Supplementary information
Supplementary Figures
Supplementary Figures 1–7
Rights and permissions
About this article
Cite this article
Karamitros, D., Stoilova, B., Aboukhalil, Z. et al. Single-cell analysis reveals the continuum of human lympho-myeloid progenitor cells. Nat Immunol 19, 85–97 (2018). https://doi.org/10.1038/s41590-017-0001-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-017-0001-2
This article is cited by
-
Loss of hematopoietic progenitors heterogeneity is an adverse prognostic factor in lower-risk myelodysplastic neoplasms
Leukemia (2024)
-
Hematopoietic stem cells with granulo-monocytic differentiation state overcome venetoclax sensitivity in patients with myelodysplastic syndromes
Nature Communications (2024)
-
Single-nucleus RNA sequencing and deep tissue proteomics reveal distinct tumour microenvironment in stage-I and II cervical cancer
Journal of Experimental & Clinical Cancer Research (2023)
-
Acute myeloid leukemia: from NGS, through scRNA-seq, to CAR-T. dissect cancer heterogeneity and tailor the treatment
Journal of Experimental & Clinical Cancer Research (2023)
-
Single-cell RNA sequencing to track novel perspectives in HSC heterogeneity
Stem Cell Research & Therapy (2022)