Abstract
Clinical prediction models in sports medicine that utilize regression or machine learning techniques have become more widely published, used, and disseminated. However, these models are typically characterized by poor methodology and incomplete reporting, and an inadequate evaluation of performance, leading to unreliable predictions and weak clinical utility within their intended sport population. Before implementation in practice, models require a thorough evaluation. Strong replicable methods and transparency reporting allow practitioners and researchers to make independent judgments as to the model’s validity, performance, clinical usefulness, and confidence it will do no harm. However, this is not reflected in the sports medicine literature. As shown in a recent systematic review of models for predicting sports injury models, most were typically characterized by poor methodology, incomplete reporting, and inadequate performance evaluation. Because of constraints imposed by data from individual teams, the development of accurate, reliable, and useful models is highly reliant on external validation. However, a barrier to collaboration is a desire to maintain a competitive advantage; a team’s proprietary information is often perceived as high value, and so these ‘trade secrets’ are frequently guarded. These ‘trade secrets’ also apply to commercially available models, as developers are unwilling to share proprietary (and potentially profitable) development and validation information. In this Current Opinion, we: (1) argue that open science is essential for improving sport prediction models and (2) critically examine sport prediction models for open science practices.
Similar content being viewed by others
References
Bullock GS, Mylott J, Hughes T, Nicholson KF, Riley RD, Collins GS. Just how confident can we be in predicting sports injuries? A systematic review of the methodological conduct and performance of existing musculoskeletal injury prediction models in sport. Sport Med. 2022;42(1):2469–82.
Bullock GS, Hughes T, Sergeant JC, Callaghan MJ, Riley RD, Collins GS. Clinical prediction models in sports medicine: a guide for clinicians and researchers. J Orthop Sport Phys Ther. 2021;51(10):517–25.
Bullock GS, Hughes T, Arundale AH, Ward P, Collins GS, Kluzek S. Black box prediction methods in sports medicine deserve a red card for reckless practice: a change of tactics is needed to advance athlete care. Sport Med. 2022;52(8):2799–801.
Bullock GS, Hughes T, Sergeant JC, Callaghan MJ, Riley R, Collins G. Methods matter: clinical prediction models will benefit sports medicine practice, but only if they are properly developed and validated. Br J Sport Med. 2021;22:1319–21.
Davis SE, Lasko TA, Chen G, Siew ED, Matheny ME. Calibration drift in regression and machine learning models for acute kidney injury. J Am Med Inform Assoc. 2017;24(6):1052–61.
Collins GS, Reitsma JB, Altman DG, Moons KG. Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD): the TRIPOD statement. J Br Surg. 2015;102(3):148–58.
Hughes T, Riley RD, Callaghan MJ, Sergeant JC. The value of Preseason screening for injury prediction: the development and internal validation of a multivariable prognostic model to predict indirect muscle injury risk in elite football (soccer) players. Sport Med Open. 2020;6(1):1–13.
Ford RA. Trade secrets and information security in the age of sports analytics. Oxford: The Oxford Handbook of American Sports Law; 2018.
Bahr R, Holme I. Risk factors for sports injuries: a methodological approach. Br J Sport Med. 2003;37(5):384–92.
Impellizzeri FM, Ward P, Coutts AJ, Bornn L, McCall A. Training load and injury part 2: questionable research practices hijack the truth and mislead well-intentioned clinicians. J Orthop Sport Phys Ther. 2020;50(10):577–84.
Caldwell AR, Vigotsky AD, Tenan MS, Radel R, Mellor DT, Kreutzer A, et al. Moving sport and exercise science forward: a call for the adoption of more transparent research practices. Sport Med. 2020;50(3):449–59.
Andrade C. HARKing, cherry-picking, p-hacking, fishing expeditions, and data dredging and mining as questionable research practices. J Clin Psychiatry. 2021;82(1):25941.
John LK, Loewenstein G, Prelec D. Measuring the prevalence of questionable research practices with incentives for truth telling. Psych Sci. 2012;23(5):524–32.
Bullock GS, Ward P, Peters S, Arundale AJH, Murray A, Impellizzeri FM, et al. Call for open science in sports medicine. Br J Sport Med. 2022;56:105719.
Wilkinson MD, Dumontier M, Aalbersberg IJ, Appleton G, Axton M, Baak A, et al. The FAIR guiding principles for scientific data management and stewardship. Sci Data. 2016;3(1):1–9.
Rockhold F, Nisen P, Freeman A. Data sharing at a crossroads. N Engl J Med. 2016;375(12):1115–7.
Bertagnolli MM, Sartor O, Chabner BA, Rothenberg ML, Khozin S, Hugh-Jones C, et al. Advantages of a truly open-access data-sharing model. N Eng J Med. 2017;376(12):1178–81.
Kadakia KT, Beckman AL, Ross JS, Krumholz HM. Leveraging open science to accelerate research. N Engl J Med. 2021;384(17): e61.
Janssen K, Moons K, Kalkman C, Grobbee D, Vergouwe Y. Updating methods improved the performance of a clinical prediction model in new patients. J Clin Epidemiol. 2008;61(1):76–86.
Riley RD, Ensor J, Snell KI, Debray TP, Altman DG, Moons KG, et al. External validation of clinical prediction models using big datasets from e-health records or IPD meta-analysis: opportunities and challenges. BMJ. 2016;353: i3140.
Riley RD, Lambert PC, Abo-Zaid G. Meta-analysis of individual participant data: rationale, conduct, and reporting. BMJ. 2010;340: c221.
Bleeker S, Moll H, Steyerberg EA, Donders A, Derksen-Lubsen G, Grobbee D, et al. External validation is necessary in prediction research: a clinical example. J Clin Epidemiol. 2003;56(9):826–32.
Tedersoo L, Küngas R, Oras E, Köster K, Eenmaa H, Leijen Ä, et al. Data sharing practices and data availability upon request differ across scientific disciplines. Sci Data. 2021;8(1):1–11.
Krawczyk M, Reuben E. (Un) available upon request: field experiment on researchers’ willingness to share supplementary materials. Acc Res. 2012;19(3):175–86.
Abalo-Núñez R, Gutiérrez-Sánchez A, Pérez MI, Vernetta-Santana M. Injury prediction in aerobic gymnastics based on anthropometric variables. Sci Sport. 2018;33(4):228–36.
Ayala F, López-Valenciano A, Martín JAG, Croix MDS, Vera-Garcia FJ, del Pilar G-V, et al. A preventive model for hamstring injuries in professional soccer: learning algorithms. Int J Sport Med. 2019;40(05):344–53.
Carbuhn AF, Sanchez Z, Fry AC, Reynolds MR, Magee LM. A simplified prediction model for lower extremity long bone stress injuries in male endurance running athletes. Clin J Sport Med. 2020;30(5):e124–6.
Carey DL, Crossley KM, Whiteley R, Mosler A, Ong K-L, Crow J, et al. Modeling training loads and injuries: the dangers of discretization. Med Sci Sport Exerc. 2018;50(11):2267–76.
Colby MJ, Dawson B, Peeling P, Heasman J, Rogalski B, Drew MK, et al. Improvement of prediction of noncontact injury in elite Australian footballers with repeated exposure to established high-risk workload scenarios. Int J Sport Physiol Perform. 2018;13(9):1130–5.
Feijen S, Struyf T, Kuppens K, Tate A, Struyf F. Prediction of shoulder pain in youth competitive swimmers: the development and internal validation of a prognostic prediction model. Am J Sport Med. 2021;49(1):154–61.
Gabbett TJ. The development and application of an injury prediction model for noncontact, soft-tissue injuries in elite collision sport athletes. J Strength Cond Res. 2010;24(10):2593–603.
Ivarsson A, Johnson U, Lindwall M, Gustafsson H, Altemyr M. Psychosocial stress as a predictor of injury in elite junior soccer: a latent growth curve analysis. J Sci Med Sport. 2014;17(4):366–70.
Jauhiainen S, Kauppi J-P, Leppänen M, Pasanen K, Parkkari J, Vasankari T, et al. New machine learning approach for detection of injury risk factors in young team sport athletes. Int J Sport Med. 2021;42(02):175–82.
Karuc J, Mišigoj-Durakovic M, Šarlija M, Markovic G, Hadžic V, Trošt-Bobic T, et al. Can injuries be predicted by functional movement screen in adolescents? The application of machine learning. J Strength Cond Res. 2021;35(4):910–9.
Khayambashi K, Ghoddosi N, Straub RK, Powers CM. Hip muscle strength predicts noncontact anterior cruciate ligament injury in male and female athletes: a prospective study. Am J Sport Med. 2016;44(2):355–61.
Landis SE, Baker RT, Seegmiller JG. Non-contact anterior cruciate ligament and lower extremity injury risk prediction using functional movement screen and knee abduction moment: an epidemiological observation of female intercollegiate athletes. Int J Sport Phys Ther. 2018;13(6):973.
López-Valenciano A, Ayala F, Puerta JM, Croix MDS, Vera-García F, Hernández-Sánchez S, et al. A preventive model for muscle injuries: a novel approach based on learning algorithms. Med Sci Sport Exerc. 2018;50(5):915.
Luu BC, Wright AL, Haeberle HS, Karnuta JM, Schickendantz MS, Makhni EC, et al. Machine learning outperforms logistic regression analysis to predict next-season NHL player injury: an analysis of 2322 players from 2007 to 2017. Orthop J Sport Med. 2020;8(9):2325967120953404.
McCann RS, Kosik KB, Terada M, Beard MQ, Buskirk GE, Gribble PA. Acute lateral ankle sprain prediction in collegiate women’s soccer players. Int J Sport Phys Ther. 2018;13(1):12.
Oliver JL, Ayala F, Croix MBDS, Lloyd RS, Myer GD, Read PJ. Using machine learning to improve our understanding of injury risk and prediction in elite male youth football players. J Sci Med Sport. 2020;23(11):1044–8.
Pontillo M, Hines SM, Sennett BJ. Prediction of ACL injuries from vertical jump kinetics in Division 1 collegiate athletes. Int J Sport Phys Ther. 2021;16(1):156.
Powers CM, Ghoddosi N, Straub RK, Khayambashi K. Hip strength as a predictor of ankle sprains in male soccer players: a prospective study. J Athl Train. 2017;52(11):1048–55.
Rommers N, Rössler R, Verhagen E, Vandecasteele F, Verstockt S, Vaeyens R, et al. A machine learning approach to assess injury risk in elite youth football players. Med Sci Sport Exerc. 2020;52(8):1745–51.
Rossi A, Pappalardo L, Cintia P, Iaia FM, Fernández J, Medina D. Effective injury forecasting in soccer with GPS training data and machine learning. PLoS One. 2018;13(7): e0201264.
Ruddy J, Shield A, Maniar N, Williams M, Duhig S, Timmins R, et al. Predictive modeling of hamstring strain injuries in elite Australian footballers. Med Sci Sport Exerc. 2018;50(5):906–14.
Shambaugh JP, Klein A, Herbert JH. Structural measures as predictors of injury basketball players. Med Sci Sport Exerc. 1991;23(5):522–7.
Sturnick DR, Vacek PM, DeSarno MJ, Gardner-Morse MG, Tourville TW, Slauterbeck JR, et al. Combined anatomic factors predicting risk of anterior cruciate ligament injury for males and females. Am J Sport Med. 2015;43(4):839–47.
Teramoto M, Cross CL, Cushman DM, Maak TG, Petron DJ, Willick SE. Game injuries in relation to game schedules in the National Basketball Association. J Sci Med Sport. 2017;20(3):230–5.
Thornton HR, Delaney JA, Duthie GM, Dascombe BJ. Importance of various training-load measures in injury incidence of professional rugby league athletes. Int J Sport Physiol Perform. 2017;12(6):819–24.
Van Der Does H, Brink M, Benjaminse A, Visscher C, Lemmink K. Jump landing characteristics predict lower extremity injuries in indoor team sports. Int J Sport Med. 2016;37(03):251–6.
Whiteside D, Martini DN, Lepley AS, Zernicke RF, Goulet GC. Predictors of ulnar collateral ligament reconstruction in Major League Baseball pitchers. Am J Sport Med. 2016;44(9):2202–9.
Wiese BW, Boone JK, Mattacola CG, McKeon PO, Uhl TL. Determination of the functional movement screen to predict musculoskeletal injury in intercollegiate athletics. Athl Train Sport Health Care. 2014;6(4):161–9.
Wilkerson GB, Colston MA. A refined prediction model for core and lower extremity sprains and strains among collegiate football players. J AThl Train. 2015;50(6):643–50.
Pollack KM, D’Angelo J, Green G, Conte S, Fealy S, Marinak C, et al. Developing and implementing Major League Baseball’s health and injury tracking system. Am J Epidemiol. 2016;183(5):490–6.
Snoke J, Raab GM, Nowok B, Dibben C, Slavkovic A. General and specific utility measures for synthetic data. J R Stat Soc A. 2018;181(3):663–88.
Abowd JM, Vilhuber L. How protective are synthetic data? International Conference on Privacy in Statistical Databases; 2008. p. 239–46.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
Gary S. Collins was supported by the NIHR Biomedical Research Centre, Oxford, and Cancer Research UK (programme Grant: C49297/A27294).
Conflict of interest
Garrett S. Bullock, Patrick Ward, Franco M. Impellizzeri, Tom Hughes, Paula Dhiman, Richard D. Riley, and Gary S. Collins have no conflicts of interest that are directly relevant to the content of this article.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Availability of data and material
Not applicable.
Code availability
Not applicable.
Authors’ contributions
GB, PW, and GSC conceived the study idea. GB, PW, FMI, TH, RR, SK, and GC were involved in the design and planning. GB and GSC wrote the first draft of the manuscript. GB, PW, FMI, SK, TH, PD, RR, and GSC critically revised the manuscript. All authors approved the final version of the manuscript.
Appendix 1: TRIPOD Checklist: Prediction Model Development and Validation
Appendix 1: TRIPOD Checklist: Prediction Model Development and Validation
Section/topic | Item | Checklist item | Page | |
|---|---|---|---|---|
Title and abstract | ||||
Title | 1 | D;V | Identify the study as developing and/or validating a multivariable prediction model, the target population, and the outcome to be predicted | |
Abstract | 2 | D;V | Provide a summary of objectives, study design, setting, participants, sample size, predictors, outcome, statistical analysis, results, and conclusions | |
Introduction | ||||
Background and objectives | 3a | D;V | Explain the medical context (including whether diagnostic or prognostic) and rationale for developing or validating the multivariable prediction model, including references to existing models | |
3b | D;V | Specify the objectives, including whether the study describes the development or validation of the model or both | ||
Methods | ||||
Source of data | 4a | D;V | Describe the study design or source of data (e.g., randomized trial, cohort, or registry data), separately for the development and validation data sets, if applicable | |
4b | D;V | Specify the key study dates, including start of accrual; end of accrual; and, if applicable, end of follow-up | ||
Participants | 5a | D;V | Specify key elements of the study setting (e.g., primary care, secondary care, general population) including number and location of centers | |
5b | D;V | Describe eligibility criteria for participants | ||
5c | D;V | Give details of treatments received, if relevant | ||
Outcome | 6a | D;V | Clearly define the outcome that is predicted by the prediction model, including how and when assessed | |
6b | D;V | Report any actions to blind assessment of the outcome to be predicted | ||
Predictors | 7a | D;V | Clearly define all predictors used in developing or validating the multivariable prediction model, including how and when they were measured | |
7b | D;V | Report any actions to blind assessment of predictors for the outcome and other predictors | ||
Sample size | 8 | D;V | Explain how the study size was arrived at | |
Missing data | 9 | D;V | Describe how missing data were handled (e.g., complete-case analysis, single imputation, multiple imputation) with details of any imputation method | |
Statistical analysis methods | 10a | D | Describe how predictors were handled in the analyses | |
10b | D | Specify type of model, all model-building procedures (including any predictor selection), and method for internal validation | ||
10c | V | For validation, describe how the predictions were calculated | ||
10d | D;V | Specify all measures used to assess model performance and, if relevant, to compare multiple models | ||
10e | V | Describe any model updating (e.g., recalibration) arising from the validation, if done | ||
Risk groups | 11 | D;V | Provide details on how risk groups were created, if done | |
Development vs validation | 12 | V | For validation, identify any differences from the development data in setting, eligibility criteria, outcome, and predictors | |
Results | ||||
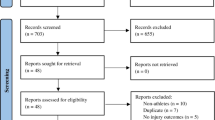
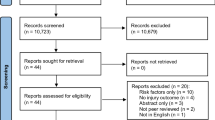
Participants | 13a | D;V | Describe the flow of participants through the study, including the number of participants with and without the outcome and, if applicable, a summary of the follow-up time. A diagram may be helpful | |
13b | D;V | Describe the characteristics of the participants (basic demographics, clinical features, available predictors), including the number of participants with missing data for predictors and outcome | ||
13c | V | For validation, show a comparison with the development data of the distribution of important variables (demographics, predictors and outcome) | ||
Model development | 14a | D | Specify the number of participants and outcome events in each analysis | |
14b | D | If done, report the unadjusted association between each candidate predictor and outcome | ||
Model specification | 15a | D | Present the full prediction model to allow predictions for individuals (i.e., all regression coefficients, and model intercept or baseline survival at a given time point) | |
15b | D | Explain how to the use the prediction model | ||
Model performance | 16 | D;V | Report performance measures (with CIs) for the prediction model | |
Model updating | 17 | V | If done, report the results from any model updating (i.e., model specification, model performance) | |
Discussion | ||||
Limitations | 18 | D;V | Discuss any limitations of the study (such as nonrepresentative sample, few events per predictor, missing data) | |
Interpretation | 19a | V | For validation, discuss the results with reference to performance in the development data, and any other validation data | |
19b | D;V | Give an overall interpretation of the results, considering objectives, limitations, results from similar studies, and other relevant evidence | ||
Implications | 20 | D;V | Discuss the potential clinical use of the model and implications for future research | |
Other information | ||||
Supplementary information | 21 | D;V | Provide information about the availability of supplementary resources, such as study protocol, Web calculator, and data sets | |
Funding | 22 | D;V | Give the source of funding and the role of the funders for the present study | |
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bullock, G.S., Ward, P., Impellizzeri, F.M. et al. The Trade Secret Taboo: Open Science Methods are Required to Improve Prediction Models in Sports Medicine and Performance. Sports Med 53, 1841–1849 (2023). https://doi.org/10.1007/s40279-023-01849-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40279-023-01849-6